To install the latest version of the Zmix package from github
library(roxygen2)
library(devtools)
install_github('zoevanhavre/Zmix')
library(Zmix)
Let's generate some data from a mixture of three Gaussian components. Using the Zmix function provided, a Gaussian mixture can be simulated from given the model parameters (\mu_k), (\sigma_k), and mixture weights (\pi_k) for (k=1,\dots, K):
simone<-simudZ(n=100,
mu=c(-1,4,10),
sig=c(1,1,2),
p=c(.2, .7, .1),
k=3)
#the output includes both the simulated data,
# and each observation's allocation as "Z".
lapply(simone, head)## $Y
## [1] 3.526882 4.880605 -1.989098 3.404140 2.320297 -1.410440
##
## $Z
## [1] 2 2 1 2 2 1
Let's run Zmix, fitting 10 components to these data.
run1 <- Zmix_univ_tempered(simone$Y,iter=500,k=10) ##
|
| | 0%
|
|============= | 20%
|
|========================== | 40%
|
|======================================= | 60%
|
|==================================================== | 80%
|
|=================================================================| 100%
 Post processing: main function
Post processing: main function
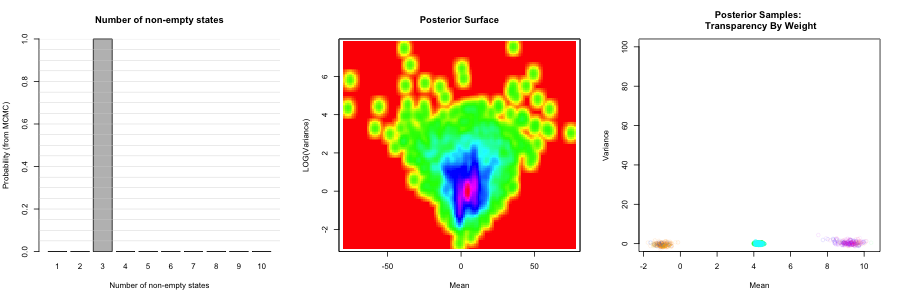
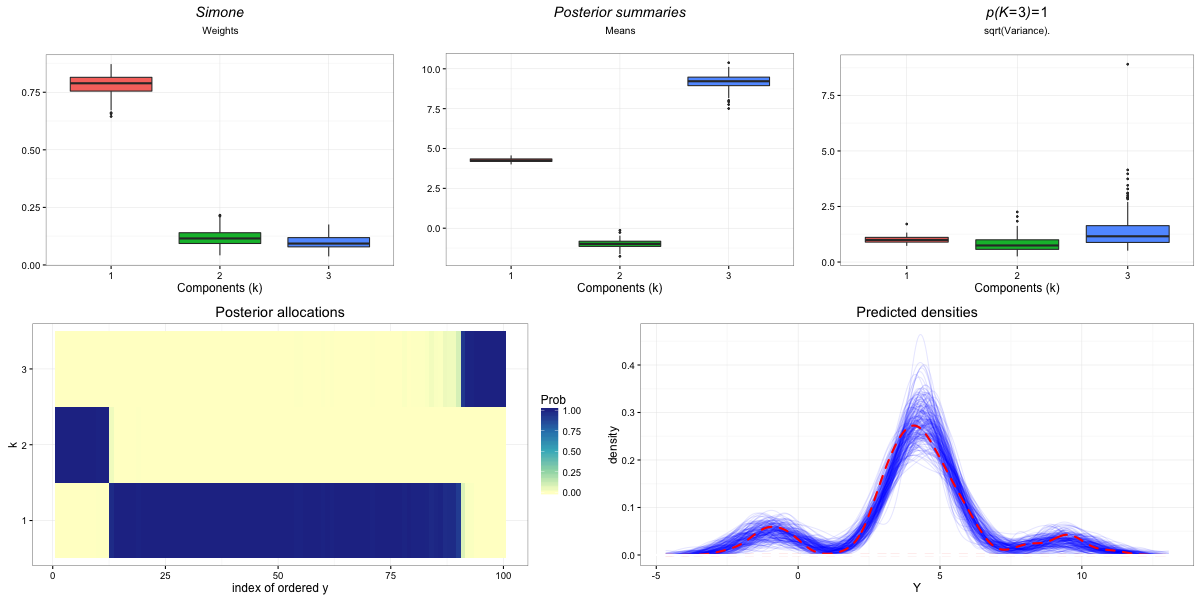
pp_run1<-Process_Output_Zmix(run1,Pred_Reps=200, Zswitch_Sensitivity=0.01, isSim=FALSE, Plot_Title="Simone", SaveFileName="Zmix_Run1", Burn=200)## NULL
## K0 Probability MAE MSE Pmin Pmax Concordance MAPE MSPE
## 1 3 1 83.86598 113.2284 0.12 0.84 0.94425 96.54466 149.0588
pp_run1[[1]]## variable factor(k) value K0
## 1 P 1 0.69(0.57,0.79) 3
## 2 Mu 1 3.98(3.68,4.25) 3
## 3 Sig 1 1.09(0.76,1.6) 3
## 4 P 2 0.18(0.11,0.26) 3
## 5 Mu 2 -1.24(-2.04,-0.51) 3
## 6 Sig 2 1.94(0.92,4.03) 3
## 7 P 3 0.14(0.07,0.23) 3
## 8 Mu 3 9.49(8.06,10.77) 3
## 9 Sig 3 2.38(0.86,6.73) 3
pp_run1[[2]]## K0 Probability MAE MSE Pmin Pmax Concordance MAPE MSPE
## 1 3 1 83.86598 113.2284 0.12 0.84 0.94425 96.54466 149.0588
The plots this makes can be found in the working directory. This includes: