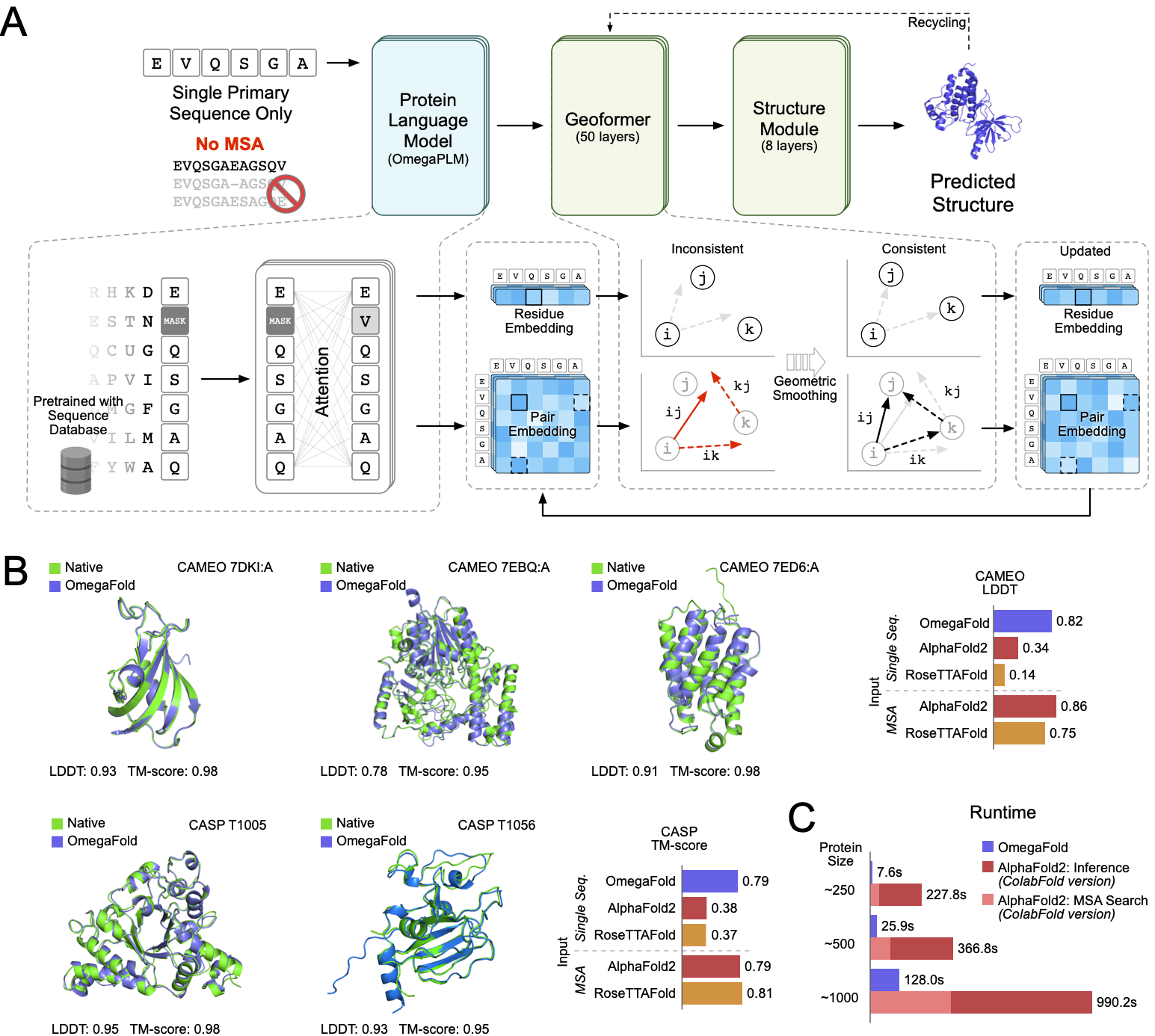
This is the first release for paper High-resolution de novo structure prediction from primary sequence.
We will continue to optimize this repository for more ease of use, for instance, reducing the GRAM required to inference long proteins and releasing possibly stronger models.
To prepare the environment to run OmegaFold,
pip install -r requirements.txt
should get you where you want. Even if this failed, since we use minimal 3rd party libraries, you can always just install the latest PyTorch and biopython (and that's it!) yourself.
There should be only one way to use the model:
python main.py INPUT_FILE.fasta OUTPUT_DIRECTORY
And voila!
The INPUT_FILE.fasta should be a normal fasta file with possibly many
sequences with a comment line starting with > or : above the amino
acid sequence itself.
This command will download the weight
from https://helixon.s3.amazonaws.com/release1.pt
to ~/.cache/omegafold_ckpt/model.pt
and load the model
However, since we have implemented sharded execution, it is possible to
- trade computation time for GRAM: by changing
--subbatch_size. The smaller this value is, the longer the execution can take, and the less memory is required, or, - trade computation time for average prediction quality, by changing
--num_cycle
For more information, run
python main.py --help
where we provide several options for both speed and weights utilities.
We produce one pdb for each of the sequences in INPUT_FILE.fasta saved in
the OUTPUT_DIRECTORY. We also put our confidence value the place of
b_factors in pdb files.
If this is helpful to you, please consider citing the paper with
@article{OmegaFold,
author = {Wu, Ruidong and Ding, Fan and Wang, Rui and Shen, Rui and Zhang, Xiwen and Luo, Shitong and Su, Chenpeng and Wu, Zuofan and Xie, Qi and Berger, Bonnie and Ma, Jianzhu and Peng, Jian},
title = {High-resolution de novo structure prediction from primary sequence},
elocation-id = {2022.07.21.500999},
year = {2022},
doi = {10.1101/2022.07.21.500999},
publisher = {Cold Spring Harbor Laboratory},
URL = {https://www.biorxiv.org/content/early/2022/07/22/2022.07.21.500999},
eprint = {https://www.biorxiv.org/content/early/2022/07/22/2022.07.21.500999.full.pdf},
journal = {bioRxiv}
}
Also some of the comments might be out-of-date as of now, and will be updated very soon