data: containsknownandunknowndatasets.eda: module that contains scripts for exploratory data analysis.models: module that contains scripts that perform parameter tuning and generate predictions for different algorithms.Genetic Algorithm(intuning.py)Random ForestLogistic RegressionSupport Vector MachinesGradient Boosted Trees
predictions: contains predictions of the various model runs - on github this is either empty or contains uninformative garbage :)preprocessing: module that contains scripts for data cleaning and feature engineering
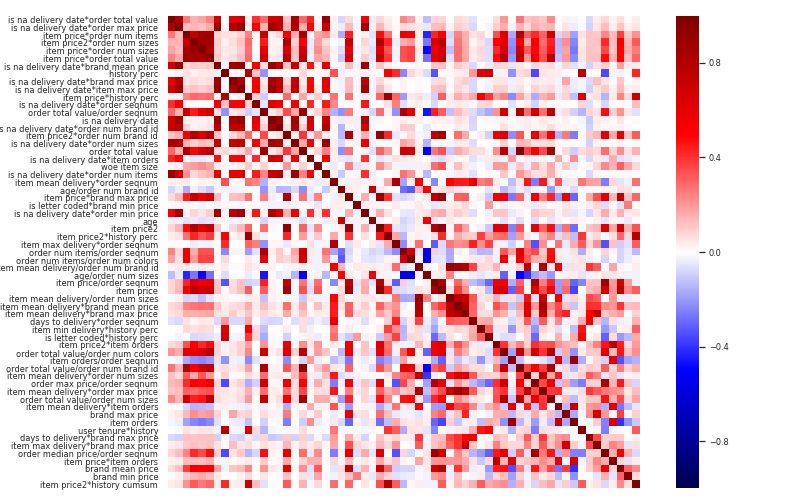
All features which had a correlation higher than 95% have been removed.
The first step is to install all requirements:
pip install -r requirements.txtThe second step is to install the repo as a package (this makes importing modules from a parent-directory possible):
pip install -e .You can skip this step. However, imports will start failing. If you do not want to install random code from an untrusted repository, you can move any notebooks/scripts you want to execute into the parent directory. This should do the trick too.
Algorithms for parameter tuning.
minimizer(objective)Create a function to minimize an objective function.
This function is intended to be used as a decorator.
objective : callable
The objective function that will be optimized. This function
should return a dictionary of the form
{"loss": loss, "status": hyperopt.STATUS_OK}.
function : callable
This decorated function will be optimized over a provided
parameter space.
sigmoid(x)Numerically stable sigmoid function.
GeneticAlgorithm(self, elitism=0.02, population_size=100, n_jobs=-1, crossover_strategy='arithmetic', prob_mutation=0.5)Minimize non-continuous functions through a Genetic Algorithm.
elitism : float
Percentage of individuals to be kept in next iteration
and used to form offspring for the next iteration.
population_size : int
Number of individuals in pool.
prob_mutation : float
Probability of mutating a particular individual.
num_parents : int
Number of parents.
n_jobs : int
Number of processors used. When -1, all processors are used, when -2
all but one are used.
history: dict
Dictionary containing information of the current run. Keeps records on
the average and optimal fitness of the population, the optimal
candidate and threshold and the out of bag fitness in each iteration.
optimal_candidate : numpy.array
Optimal parameter vector.
optimal_cutoff : float
Optimal threshold for transforming probablistic
predictions into class labels.
fit(self, X, y, price, fit_intercept=True, loc=0, scale=1) :
Fit features to data. This is especially useful to avoid data leaks.
run(self, maxiter=10, subsample=None, bootstrap=False, reset_prob=1) :
Run Genetic Algorithm, determine optimal coefficients and threshold.
predict_proba(self, X, b=None) :
Return predicted probabilities of the positive class based on the
optimal coefficients.
predict(self, X, cut=None) :
Return binary predictions based on the optimal coefficients and thres-
hold.
predict_cost(self, X=None, y_true=None, price=None, b=None, cutoff=None) :
Return predicted cost.
get_fitness(self, y_prob, y_true, price) :
Compute optimal costs of misclassification and optimal threshold.
get_utility(self, y_prob=None, y_true=None, price=None, cutoff=None) :
Compute costs of misclassification based on arbitrary threshold.
plot(self, savepath=None, title=None, **kwargs) :
Draw diagnostic plot.
GeneticAlgorithm.fit(self, X, y, price, fit_intercept=True, loc=0, scale=1)Stage Genetic Algorithm with regards to data.
X : numpy.ndarray
Train-data to calculate fitness with.
y : array-like
Vector of true labels.
price : array-like
Vector of item prices.
fit_intercept : bool
Flag if intercept should be fitted.
loc : float
Location parameter during initialization.
scale : float
Scale parameter during initialization.
self
GeneticAlgorithm.run(self, maxiter=10, subsample=None, bootstrap=False, reset_prob=1)Train the algorithm.
maxiter : int
Maximum number of training iterations.
subsample : float
Percentage of training data to be used for creating a subsample.
bootstrap : float
If true, draw with replacement.
reset_prob : float
Chance of redrawing the sample.
optimal_candidate : numpy.array
GeneticAlgorithm.plot(self, savepath=None, title=None, **kwargs)Create diagnostic plots
savepath : str
If set, save plot to savepath.
title : str
Title of figure. If None (default) omit title.
kwargs : dict
Additional arguments to pyplot.subplots.
GeneticAlgorithm.predict_proba(self, X, b=None)Calculate predicted probabilities of positive class.
X : numpy.ndarray
Feature matrix.
b : numpy.array
Parameter vector. If None (default) use optimal parameter
of the last iteration.
out : numpy.array
Probabilities of the positive class computed by a logit-model.
GeneticAlgorithm.predict(self, X, cut=None)Calculate predicted class labels.
X : numpy.ndarray
Feature matrix.
cut : float
Threshold for prediction. If probablity is larger than thres-
hold predict positive class. If None (default) use the opt-
imal parameter of the last iteration.
out : numpy.array
Predicted class labels.
GeneticAlgorithm.predict_cost(self, X=None, y_true=None, price=None, b=None, cutoff=None)Return predicted cost.
Predict cost for a given feature matrix, labels, price and cutoff or calculate costs on the training set based on the optimal solution.
X : numpy.ndarray
Feature matrix. If None (default) use the whole training data.
y_true : array-like
Vector of true labels. If None (default) use the whole
training data.
price : array-like
Vector of prices. If None (default) use the whole training
data.
b : array-like
Vector of coefficients. If None use the optimal solution of the
last iteration.
cutoff : float
Threshold for prediction. If None (default) use the optimal
threshold of the last iteration.
out : float
Predicted cost.
GeneticAlgorithm.get_fitness(self, y_prob, y_true, price)Calculate optimal fitness and threshold.
Evaluate cost/utility function on a grid of thresholds and return the optimal values.
y_prob : array-like
Vector of predicted probablities of positive class.
y_true : array-like
Vector of true labels.
price : array-like
Vector of item prices.
best_util : float
Best achievable utility.
best_cut : float
Threshold that leads to optimal utility.
GeneticAlgorithm.get_utility(self, y_prob=None, y_true=None, price=None, cutoff=None)Calculate cost/utility.
Evaluate based on predicted probabilities and a threshold.
y_prob : array-like
Vector of predicted probabilities of positive class.
y_true : array-like
Vector of true labels.
price : array-like
Vector of item prices.
cutoff : float
Threshold for creating predictions.
utility : float
Cost/utility.
Clean the dataset.
Gather all functions which clean the data and compute features that do not require aggregation.
clean(datapath)Clean BADS data.
Read the data and perform the following tasks:
- parse dates:
order_date,delivery_date,user_dob,user_reg_date. - replace non-standard coding for missing values.
- drop esoteric colors.
- impute missing values for
delivery_datecolumn by its mean. - remove odd reported values for user dates of birth.
- impute missing
user_dob:user_reg_date - mean(user_reg_date-user_dob) - extract dummies according to
sizecolumn:is_item_pants: ifsizematches exactly 4 numbers.is_item_clothes: ifsizeis notunsized.is_item_underwear: ifsizematches up to 2 numbers.is_letter_coded: ifsizenotunsizedbut contains characters.
- extract features according to
delivery_date:delivery_thu: indicator if item was delivered on a Thursday.delivery_fri: indicator if item was delivered on a Friday.days_to_delivery: number of days between order and delivery.
datapath : str Path to dataset.
data : pd.DataFrame Cleaned dataset.
FeatureGenerator(self, cols=None)Generate Features for the different algorithms.
This class interface facilitates the fitting and transforming of different datasets and gathers the numerous aggregation levels.
cols : list
List of column names to be returned at most. Must be a subset of all
columns that are defined within the fit method.
cols : list
Column names of the returned array.
dropcols : list
Column names that should be dropped, because their correlation
with other variables is too high.
target : pd.Series
Series of class values/labels.
features : pd.DataFrame
Original features, copy of data.
item_woe : pd.DataFrame
item_id and associated weight of evidence.
color_woe : pd.DataFrame
item_color and associated weight of evidence.
brand_woe : pd.DataFrame
brand_id and associated weight of evidence.
size_woe : pd.DataFrame
item_size and associated weight of evidence.
items : pd.DataFrame
Contains general information on items. Groupby-Keys: item_id.
orders : pd.DataFrame
Contains general information on orders. Groupby-Keys: user_id
order_date.
brands : pd.DataFrame
Contains general information on brands. Groupby-Keys: brand_id.
states : pd.DataFrame
Contains general information on states. Groupby-Keys: user_state.
outfeatures : pd.DataFrame
DataFrame version of the returned numpy.ndarray.
fit(self, data, target_col) :
Fit data.
transformself, X, ignore_woe=True, add_dummies=False, add_interactions=True, add_ratios=True) :
Tranform data.
fit_transform(self, data, target_col, ignore_woe=True, add_dummies=False, add_interactions=True, add_ratios=True) :
Fit data, then transform it.
FeatureGenerator.fit(self, data, target_col)Compute aggregated data according to different levels.
The different aggregation levels combine information on:
- items: group by
item_id - orders: group by
user_idandorder_date - brands: group by
brand_id - states: group by
states
data : pd.DataFrame
Full set of information to be used. This can be the known
dataset or a concatenation of known and unknown.
target_col : str
Name of the column that contains the labels
FeatureGenerator.transform(self, X, ignore_woe=True, add_dummies=False, add_interactions=True, add_ratios=True)Add aggregated information to X and compute additional features.
X : pd.DataFrame
DataFrame to be transformed.
ignore_woe : bool, optional
Flag if Weight of Evidence columns should be ignored.
Default is True.
out, [y] : np.ndarray
Note that y will only be returned if columns of X
contain the target column.
FeatureGenerator.fit_transform(self, data, target_col, ignore_woe=True, add_dummies=False, add_interactions=True, add_ratios=True)Fit data then transform it.