A collaborate platform for streamlining a BIDS workflow for the NENAH study, on the linux workstation based at University of Southampton.
This project is to streamline a Brain Imaging Data Structure (BIDS) workflow for Neurodevelopmental Trajectories and Neural Correlates in Children with Neonatal Hypoxic-Ischaemic Encephalopathy (NENAH), i.e. to put the NENAH data into a BIDS data structure, for Quality Assessment and customisable preprocessing, so as to keep track of the data quality continuously and quickly decide on recalls. Potentially, NeuroImaging Data Model (NIDM) can be applied to turn BIDS datasets into Semantic-BIDS datasets with relevant clinical data, e.g. test scores.
All the tasks are organized into different Projects of this repository. Specifically:
- BIDS data structure tracks the process of converting the raw DICOMs into a BIDS structured data set with relevant Metadata.
- Software management tracks study-specific software on the linux workstation.
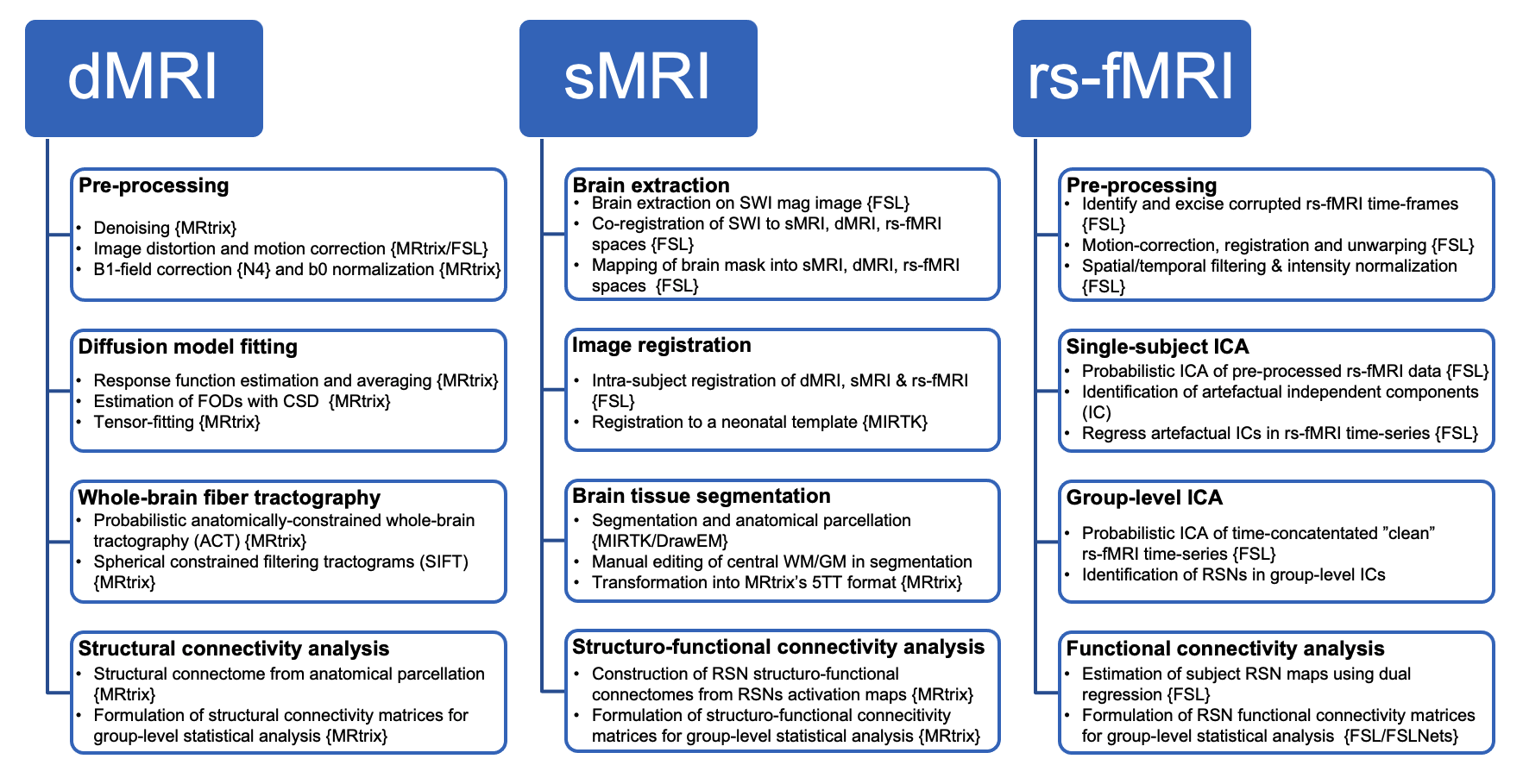
- Preprocessing of specific modalities will be tracked using dMRI pipeline and fMRI pipeline.
BIDS conversion is set up using heudiconv (a heuristic-centric DICOM converter), mimicking the tutorial here.
For quality assessment of sMRI and rs-fMRI, MRIQC is used. For dMRI, the QC outputs from FSL eddy or eddy_quad will be used.
- BIDS and the NeuroImaging Data Model (NIDM)
- dHCP: Data Release 2021, Data Release 2019, Structural Pipeline; Quality Control
- ABCD-ReproNim Course (syllabus): NIDM at Week 6 - 6th November, 2020
- Neurostars: forum for BIDS and ABCD-ReproNim discussions
- Winawer Lab [Sample Data Pipeline]
- Iridis: Singularity and Docker, also watch ABCD-ReproNim Week 3 videos
- QSIprep: Preprocessing and analysis of q-space images
- Neuroimaging and Data Science