SeqSizzle is a pager for viewing FASTQ files with fuzzy matching, allowing different adaptors to be colored differently.
Latest pre-built release binary
./SeqSizzle -h:
Usage: SeqSizzle [OPTIONS] <FILE>
Arguments:
<FILE> The FASTQ file to view
Options:
--adapter-3p
Start with 10x 3' kit adaptors:
- Patrial Read1: CTACACGACGCTCTTCCGATCT (and reverse complement)
- Partial TSO: AGATCGGAAGAGCGTCGTGTAG (and reverse complement)
- Poly(>10)A/T
--adapter-5p
Start with 10x 5' kit adaptors
- Patrial Read1: CTACACGACGCTCTTCCGATCT (and reverse complement)
- TSO: TTTCTTATATGGG (and reverse complement)
- Patrial Read2: AGATCGGAAGAGCACACGTCTGAA (and reverse complement)
- Poly(>10)A/T
-p, --patterns <PATTERNS_PATH>
Start with patterns from a CSV file
Must have the following header:
pattern,color,editdistance,comment
-s, --save-patterns <SAVE_PATTERNS_PATH>
Save the search panel to a CSV file before quitting.
To be removed in the future since you can now hit Ctrl-S in
the search panel to save the patterns
-h, --help
Print help
-V, --version
Print version
 Up / down arrow (or
Up / down arrow (or j / k) to scroll by one line, Ctrl+U / Ctrl+D to scoll half a screen.
/ (or Ctrl+F) to toggle search panel, q to quit
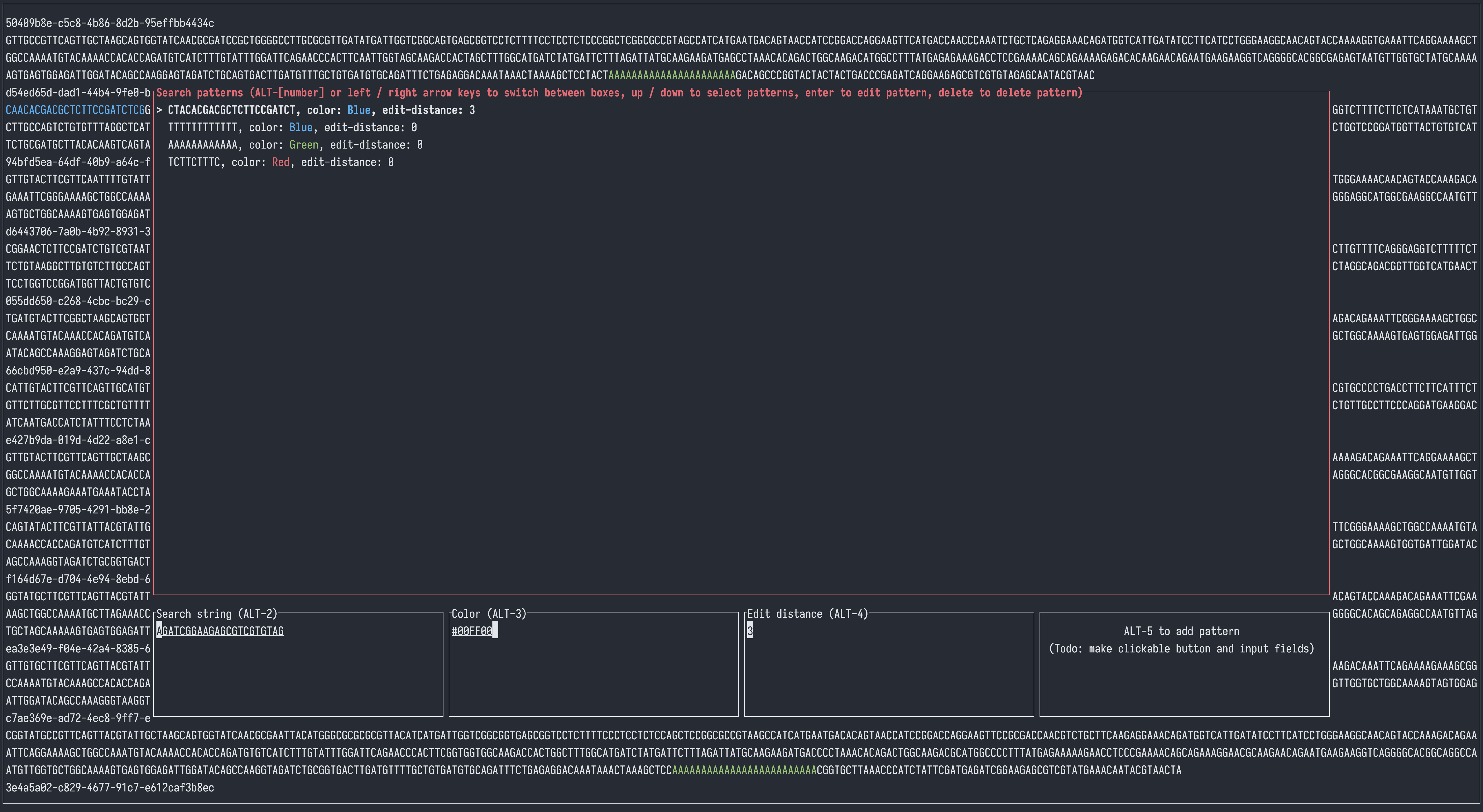 Left / right arrow (or Tab / Shift-Tab) to cycle through different input fields and the patterns list.
Left / right arrow (or Tab / Shift-Tab) to cycle through different input fields and the patterns list.
When on the patterns list field, up / down arrows cycle through patterns, Backspace (or Delete, d) to delete the selected pattern and Return to pop the pattern into the input fields for editing.
Return to add current inputs into the search pattern list (when focusing on any of the input boxes, rather than the patterns list).
Use Shift + arrow keys to move cursor within an input field (as arrow keys alone are bind to cycling input fields).
/ or Esc to close the search panel.
- Gzip (
fastq.gz) support - Filter reads by match
- Counting reads with match
- Make elements in the search panel clickable, try implementations discussed in ratatui repo
- Unit tests