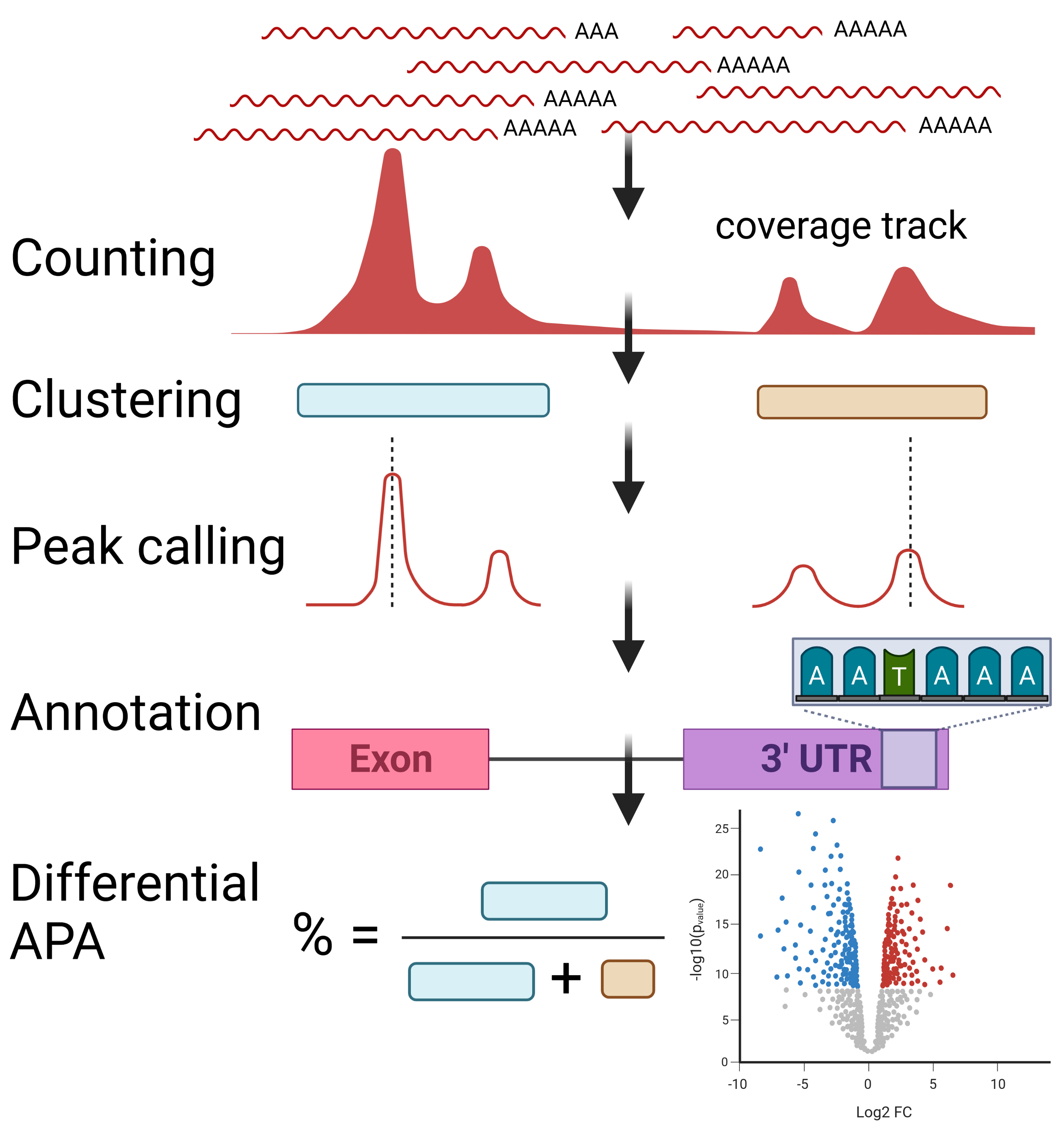
Alternative polyadenylation detection from diverse data sources such as 3'-seq, long-read and short-reads.
pip install lapa
lapa --alignment {rep1.bam},{rep2.bam},{rep3.bam} \
--fasta {fasta} \
--annotation {gtf} \
--chrom_sizes {chrom_sizes} \
--output_dir {output}
Argument details (doc):
$ lapa --help
Usage: lapa [OPTIONS]
CLI interface for lapa polyA cluster calling.
Options:
--alignment TEXT Single or multiple bam file paths are
separated with a comma.Alternatively, CSV
file with columns of sample, dataset, path
where the sample columns contains the name
of the sample, the dataset is the group of
samples replicates of each other, and path
is the path of bam file. [required]
--fasta TEXT Genome reference (GENCODE or ENSEMBL fasta)
[required]
--annotation TEXT Standart genome annotation (GENCODE or
ENSEMBL gtf). GENCODE gtf file do not
contains annotation for `five_prime_utr` and
`three_prime_utr` so need to be corrected
with `gencode_utr_fix` (see https://github.c
om/MuhammedHasan/gencode_utr_fix.git).
[required]
--chrom_sizes TEXT Chrom sizes files (can be generated with
`faidx fasta -i chromsizes > chrom_sizes`)
[required]
--output_dir TEXT Output directory of LAPA. See
lapa.readthedocs.io/en/latest/output.html)
for the details of the directory structure
and file format. [required]
...
Recommend setting is including all the samples with thier biosample/experimental replicates (tissue, cell line):
samples.csv
sample,dataset,path
ENCFF772LYG,myoblast,ENCFF772LYG.bam
ENCFF421MIL,myoblast,ENCFF421MIL.bam
ENCFF699KOR,myotube,ENCFF699KOR.bam
ENCFF731HHB,myotube,ENCFF731HHB.bam
then LAPA takes samples_config.csv as input:
lapa --alignment samples.csv \
--fasta {fasta} \
--annotation {gtf} \
--chrom_sizes {chrom_sizes} \
--output_dir {output}
...
lapa_tss --alignment samples.csv \
--fasta {fasta} \
--annotation {gtf} \
--chrom_sizes {chrom_sizes} \
--output_dir {output}
Argument details (doc):
$ lapa_tss --help
Usage: lapa_tss [OPTIONS]
CLI interface for lapa tss cluster calling.
Options:
--alignment TEXT Single or multiple bam file paths are
separated with a comma.Alternatively, CSV
file with columns of sample, dataset, path
where the sample columns contains the name
of the sample, the dataset is the group of
samples replicates of each other, and path
is the path of bam file. [required]
--fasta TEXT Genome reference (GENCODE or ENSEMBL fasta)
[required]
--annotation TEXT Standart genome annotation (GENCODE or
ENSEMBL gtf). GENCODE gtf file do not
contains annotation for `five_prime_utr` and
`three_prime_utr` so need to be corrected
with `gencode_utr_fix` (see https://github.c
om/MuhammedHasan/gencode_utr_fix.git)
[required]
--chrom_sizes TEXT Chrom sizes files (can be generated
with)`faidx fasta -i chromsizes >
chrom_sizes`) [required]
--output_dir TEXT Output directory of LAPA. See
lapa.readthedocs.io/en/latest/output.html)
for the details of the directory structure
and file format. [required]
See the following documentation links for other features, parameters of LAPA, python api and statistical testing:
Readthedocs: https://lapa.readthedocs.io/en/latest/index.html
API reference: https://lapa.readthedocs.io/en/latest/autoapi/index.html
Colab tutorials (analysis of myoblast myotube cell differentiation): https://colab.research.google.com/drive/1QzMxCRjCk3i5_MuHzjozSRWMaJgdEdSI?usp=sharing
If you are using LAPA on academic studies cite the following paper:
@article{celik2022analysis,
title={Analysis of alternative polyadenylation from long-read or short-read RNA-seq with LAPA},
author={Celik, Muhammed Hasan and Mortazavi, Ali},
journal={bioRxiv},
year={2022},
publisher={Cold Spring Harbor Laboratory}
}