This software package provides Matlab implementations of a number of algorithms for removing cardiac interference from surface EMG measurements, as well as metrics and two exemplary datasets for evaluating their respective accuracy.
In particular, all algorithms desribed in the following two publications are included:
- E. Petersen, J. Sauer, J. Grasshoff, P. Rostalski, "Removing Cardiac Artifacts from Single-Channel Respiratory Electromyograms", IEEE Access, 2020, and
- E. Petersen, "Model-based Probabilistic Inference for Monitoring Respiratory Effort using the Surface Electromyogram", Dissertation, forthcoming.
In case of any questions, comments, or bugs, please don't hesitate to contact me or open an issue on GitHub.
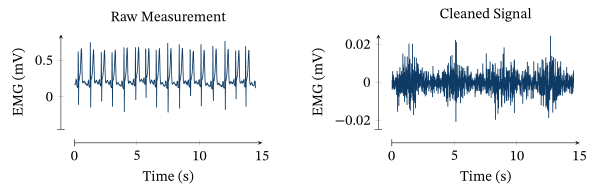
The following picture shows an example of the problem to be solved here (and the solution provided by the PATS algorithm, see below):
Two exemplary data sets of respiratory electromograms (two channels repectively) and simultanously measured airway pressure.
Matlab implementations of following algorithms:
- EMG Preprocessing
- QRS Detection (mostly taken from the OSET toolbox of Reza Sameni, with some minor modifications)
- Template Subtraction and Adaptive Template Subtraction
- Model-based ECG removal with second and 25th order Extended Kalman filter and smoother (based on Sameni et al., "A nonlinear Bayesian filtering framework for ECG denoising," 2007, IEEE Transactions on Biomedical Engineering, and Akhbari et al., "ECG denoising and fiducial point extraction using an extended Kalman filtering framework with linear and nonlinear phase observations," Physiological Measurement, 2016). Implementation partially based on the OSET toolbox of Reza Sameni.
- Wavelet Denoising (contributed by Jan Graßhoff)
- Empirical Mode Decomposition
- Probabilistic Adaptive Template Subtraction (PATS, best-performing algorithm as per Petersen 2022).
Performance evaluation functions
- Signal-to-noise ratio
- Periodicity Measure
- Power spectral density
Analysis script: ecg_removal.m and functions to read, filter, plot and evaluate data
ecg_removal.m
to read the provided data sets, apply ECG-removal algorithms (or reload precomputed results) and execute a performance evaluation.
The toolbox contains code contributions by Reza Sameni (Pan-Tompkins peak detection and some of the files in the model-based directory - see individual file headers -, all from the OSET toolbox), Jan Graßhoff (swt_denoising) and Julia Sauer (various files throughout the project).
Julia Sauer also performed the study from which the two included recordings have been obtained.
Everything else is authored by me.
Most of the work leading to the development of this toolbox has been done while I was at the University of Lübeck, with the Institute for Electrical Engineering in Medicine.
Eike Petersen, 2021