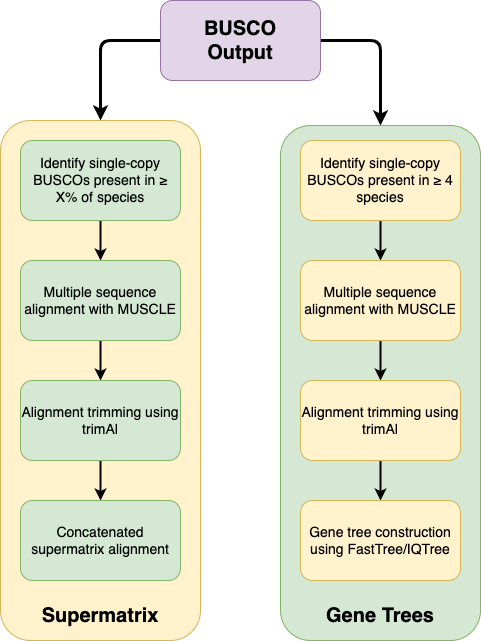
This is a Python pipeline to construct species phylogenies using BUSCO proteins. It works directly from BUSCO output and can generate concatenated supermatrix alignments and also gene trees of BUSCO families.
The pipeline identifies BUSCO proteins that are complete and single-copy in all input samples. Alternatively, you can account for missing data and choose to include BUSCO proteins that are complete and single-copy in a certain percentage of input samples. Each BUSCO family is individually aligned, trimmed, and then concatenated together to generate a supermatrix alignment. The pipeline also identifies BUSCO proteins that are complete and single-copy in at least 4 input samples, and generates gene trees for each of these families.
The pipeline requires the following dependencies:
These should be available from your $PATH. Alternatively, they can be installed using conda with the provided yaml file conda_env.yaml, which will create a conda environment called BUSCO_phylogenomics
git clone https://github.com/jamiemcg/BUSCO_phylogenomics
cd BUSCO_phylogenomics
conda env create -f conda_env.yaml
conda activate BUSCO_phylogenomics
python BUSCO_phylogenomics.py --help
usage: BUSCO_phylogenomics.py [-h] -i INPUT -o OUTPUT -t THREADS [--supermatrix_only] [--gene_trees_only] [-psc PSC] [--trimal_strategy TRIMAL_STRATEGY] [--missing_character MISSING_CHARACTER] [--gene_tree_program GENE_TREE_PROGRAM] [--busco_version_3]
Perform phylogenomic reconstruction using BUSCO proteins
options:
-h, --help show this help message and exit
-i INPUT, --input INPUT
Input directory containing completed BUSCO runs
-o OUTPUT, --output OUTPUT
Output directory to store results
-t THREADS, --threads THREADS
Number of threads to use
--supermatrix_only Don't generate gene trees
--gene_trees_only Don't perform supermatrix analysis
--nt Align nucleotide sequences instead of amino acid
sequences
-psc PSC, --percent_single_copy PSC
BUSCO presence cut-off. BUSCOs that are complete and
single-copy in at least [-psc] percent of species will
be included in the contatenated alignment
[default=100.0]
--trimal_strategy TRIMAL_STRATEGY
trimal trimming strategy (automated1, gappyout,
strict, strictplus) [default=automated1]
--missing_character MISSING_CHARACTER
Character to represent missing data [default='?']
--gene_tree_program GENE_TREE_PROGRAM
Program to use to generate gene trees (fasttree or
iqtree) [default=fasttree]
--busco_version_3 Flag to indicate that BUSCO version 3 was used (which
has slighly different output structure)
You should move all of your completed BUSCO output directories into the same directory.
Example usage:
python BUSCO_phylogenomics.py -i BUSCO_results -o output_busco_phylogenomics -t 8
This will look in the "BUSCO_results" directory for completed BUSCO runs, generate multiple sequence alignments for all complete single-copy proteins that were found in all samples, trim alignments with trimal and then concatenate them together, generating a concatenated alignment in Fasta and Phylip format along with a partitions file in NEXUS format. It will also generate gene trees for all BUSCO proteins that are complete and single-copy in at least 4 samples. The output will be stored in a directory named "output_busco_phylogenomics". The pipeline is written to be executed on a single node/machine, here 8 parallel alignment/trimming/phylogeny jobs would run.
If you don't want to generate gene trees, you can use the parameter --supermatrix-only to only generate the concatenated alignment.
If you don't want to generate a concatenated alignment, you can use the parameter --gene_trees_only to only generate gene trees.
By default, the pipeline works in protein space (i.e., aligns amino acid sequences). The --nt flag switches to using BUSCO nucleotide sequences instead of proteins.
If you have a patchy dataset and want to include BUSCO proteins in your concatenated alignment that aren't universally present, you can use the --percent_single_copy parameter.
For example:
python BUSCO_phylogenomics.py -i BUSCO_results -o output_busco_phylogenomics -t 8 --percent_single_copy 70
will include all BUSCO families that are complete and single-copy in at least 70% of samples in your concatenated alignment. Missing data will be represented by "?" characters in the concatenated alignment by default. You can specify a different character to represent missing data with the --missing_character parameter.
The provided count_buscos.py script can be used to count single-copy BUSCOs and summarise BUSCO presence/absences across
samples to determine an appropriate cut-off for how much missing data to allow (--percent_single_copy).
python count_buscos.py -i BUSCO_runs
This will report how many BUSCOs are complete and single-copy in what percentage of samples and print a presence/absence table for each BUSCO family.
If you used BUSCO version 3 you should use the flag --busco_version_3 as the output structure of this version of BUSCO is slightly different to that of versions 4 and 5.