Background
Breast cancer is an unfortunate ailment that affects millions of people worldwide. Personally, it has touched many people close in my family which is what made me interested in this data. There is much research dedicated to identifying tumors early in order to conduct invasive procedures before the cancer progresses to a late stage. One particular area of interest is using machine learning to classify a tumor as malignant (cancerous) or benign. The University of Wisconsin has publicly published data on tumors for people to look into what methods are better at classifying them.
For this project, I decided to compare logistic regression and random forests, two popular methods in binary classification problems. I also decided to use k-fold cross validation to efficiently test multiple runs of each model.
Quick Data Exploration
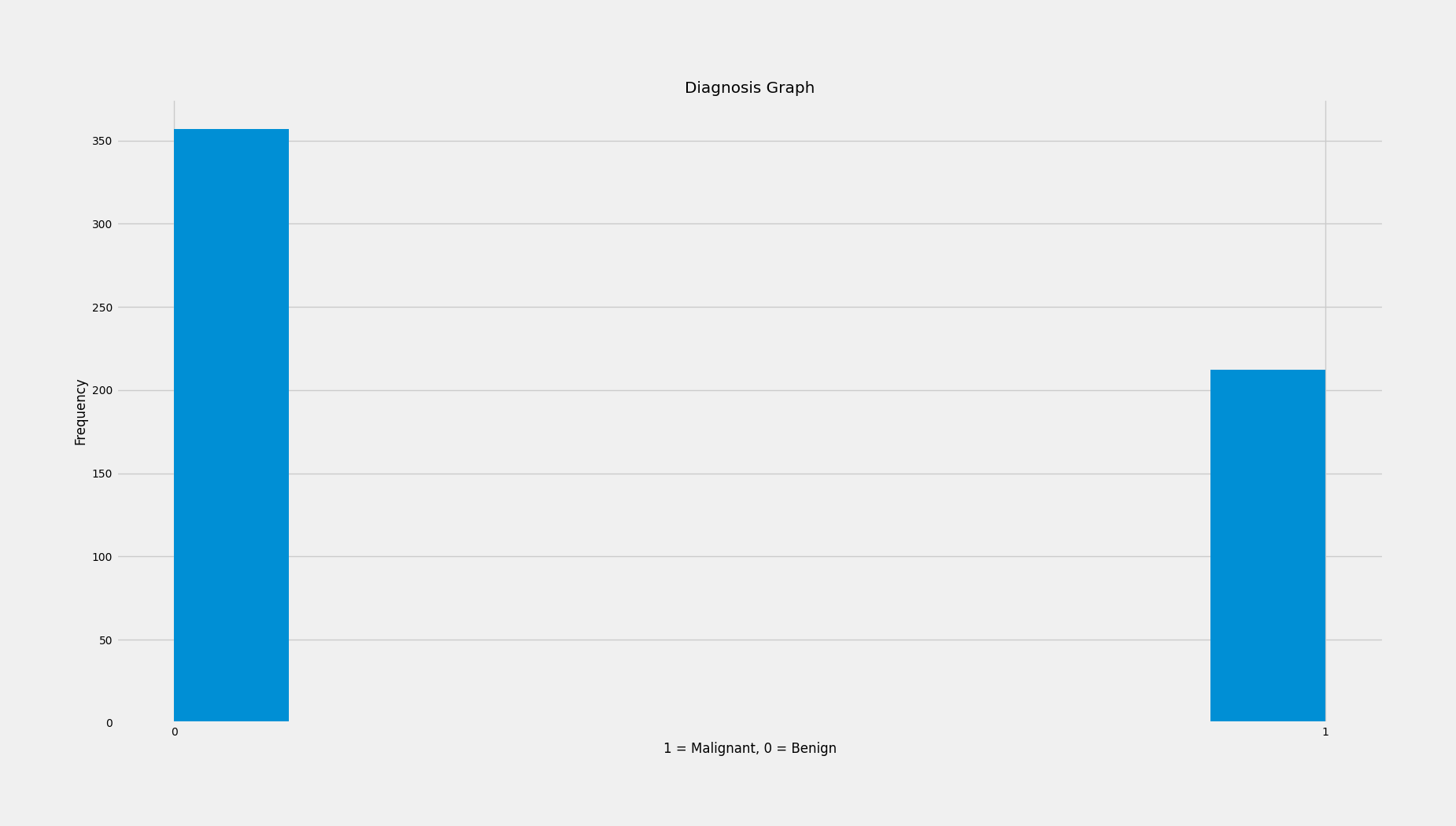
From the histogram of our dependent variable (malignant or benign) we can see that our data contains more benign tumors. Specifically we have 357 benign tumors and 212 malignant tumors.
For the independent variables, this dataset contains many! We have radius, texture, perimeter, area, smoothness, compactness, concavity, concave points, symmetry, and fractal dimension. We also have the standard error for each along with the "worst" or largest of each.
Analysis
For code, please see python file
Ok! Let's start with logistic regression with all of our variables excluding the standard error or the "worst" ones.
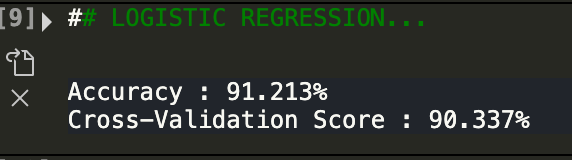
The accuracy score is running the model once. The cross validation score is the average after doing 5 folds. As we can see, logistic regression had a pretty good result! It yieled a cross validation score of about 90 percent. While this is good, let's see if random forests can do any better.
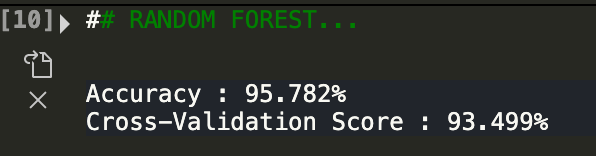
As we can see, random forests using the same set of variables did even better than logistic regression! It yielded 93% accuracy. Now let's see if we can make this even better. Let's take out all variables related to size ie. radius, perimeter, and area.
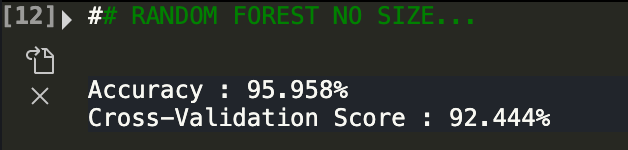
While still pretty accruate, the cross validation score went down by one percentage point. Ok, now let's look at only area variables.
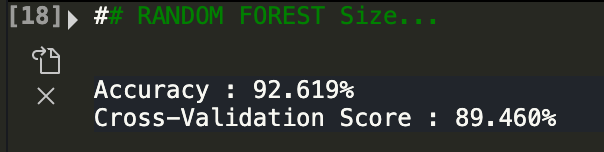
As we can see, only using size variables makes the predictions worse.
For fun, now let's see what happens if we use all variables (regular + standard error + worst of them)
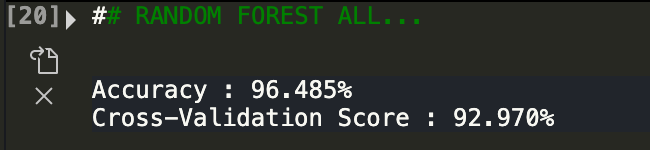
As we can see, the model performs well but not much any better than if we only use all of the regular variables! From this analysis, we can see that when assessing a tumor, a holisitc approach needs to be done. This makes sense and cancers differ from person to person. What might be a red flag in one person may not be anything in another! Hope you enjoyed reading this project. Breast cancer is something that touches many families. As people interested in data, we all have a role to play in improving analytics that help people and save lives!