huangch
This library is an extension of OpenSlide (you probably should not be here if you don't know what it is). OpenRemoteSlide is just similar to OpenSlide, except OpenRemoteSlide can read whole slide images from remote.
In OpenSlide, the tool openremoteslide-show-properties is able to show the properties of a whole slide image file:
openslide-show-properties path/to/a/wsi
E.g., given a whole slide image file 54089012-2e2f-453f-8b88-7f80d6791eb7.svs, you can extract the properties of the svs by:
./openslide-show-properties ./54089012-2e2f-453f-8b88-7f80d6791eb7.svs
In OpenRemoteSlide, you can do this:
openslide-show-properties url/to/a/wsi
E.g., you can access TCGA whole slide image (in svs format) by:
./openslide-show-properties https://api.gdc.cancer.gov/legacy/data/54089012-2e2f-453f-8b88-7f80d6791eb7
where 54089012-2e2f-453f-8b88-7f80d6791eb7 is the UUID of a TCGA whole slide image.
Here is another example: the command,
./openslide-write-png https://api.gdc.cancer.gov/legacy/data/54089012-2e2f-453f-8b88-7f80d6791eb7 0 0 3 3237 2669 output.png
helps you to read a defined region of interest on a whole slide image of TCGA over the Internet.
OpenRemoteSlide is compatible with OpenSlide (thus, compatible with OpenSlide-Python: https://github.com/openslide/openslide-python and OpenSlide-Java: https://github.com/openslide/openslide-java), except the Windows version.
Recent progress in applying deep learning technologies for the analysis of digital pathological images has received much attention from researchers in both industry and academia as accuracy of these methods can be as good as 98.4%. However, most deep learning approaches require huge sets of training patterns. In addition, a pathological whole slide image (WSI) can be as big as 1 to 10 GB and most of the images are not always freely available on a local storage system. On the other hand, not every piece on the WSI is needed for either human-made or automated pathological analysis. This fact represents a huge cost redundancy in terms of data transfer and storage.
Thus, I decided to develop OpenRemoteSlide, which is an extension for OpenSlide (https://openslide.org/), allowing users accessing images from remote based on the same fashion of using OpenSlide. For example, in OpenSlide, you use (/path/to/an/image/file.svs) pointing a digital pathological whole-slide image file. In OpenRemoteSlide, instead of downloading all giga-size image files beforehand, you can actually access any region of an image from remote by giving its URL, e.g., (https://url/to/an/image/file.svs), so that the cost for data transfer and storage can be saved.
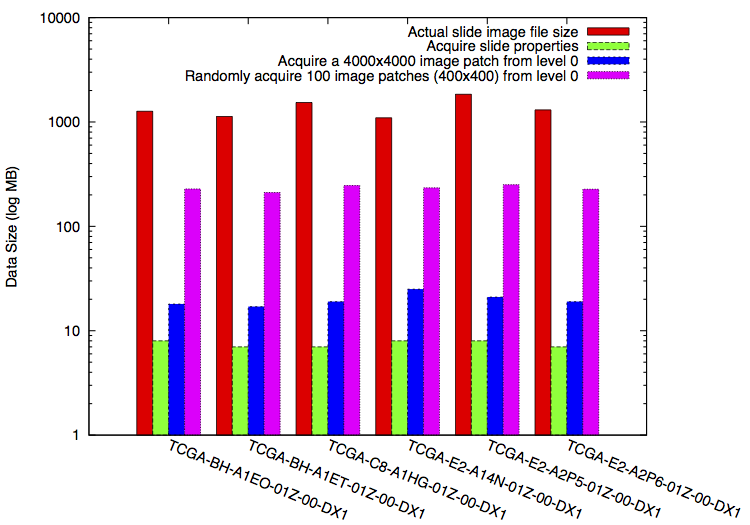
The figure shows comparison of the costs of data transfer and storage performance for OpenRemoteSlide. We used the TCGA pathological image set as an example, selected some WSIs with size ~1GB. This plot shows the differences between the costs of acquiring various amount of information from these WSIs.
Based on OpenRemoteSlide, the average cost (the actual size of data transferred and stored) for obtaining the image properties is ~7.5 MB, which is less than 1% of the data file size of the chosen image. For acquiring a (4000x4000) image from level 0, the average cost is ~19.8 MB, about 1.8% of the total data size. For randomly acquiring 100 image samples with size of (400x400), the average cost is ~233 MB, about 21.2% of the total data size. In other words, the proposed OpenRemoteSlide can save the cost of accessing WSIs from remote from 78.8% up to 99.3%, depending on the desired coefficients for the data acquisition.
For the other details, please see README-OpenSlide.txt. You can also find the original distribution of OpenSlide from: https://openslide.org
Good luck!