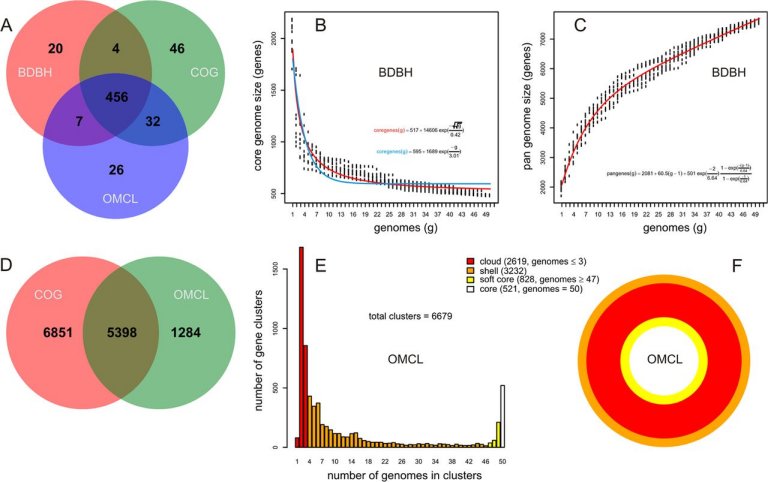
This software is maintained by Bruno Contreras-Moreira (bcontreras at eead.csic.es) and Pablo Vinuesa (vinuesa at ccg.unam.mx). The original version, suitable for bacterial genomes, was described in:
Contreras-Moreira B, Vinuesa P (2013) Appl. Environ. Microbiol. 79:7696-7701
Vinuesa P, Contreras-Moreira B (2015) Methods in Molecular Biology Volume 1231, 203-232
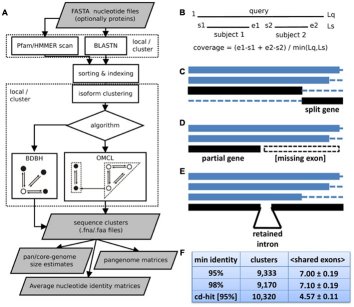
The software was subsequently adapted to the study of intra-specific eukaryotic pan-genomes resulting in script GET_HOMOLOGUES-EST, described in:
Contreras-Moreira B, Cantalapiedra CP et al (2017) Front. Plant Sci. 10.3389/fpls.2017.00184
GET_HOMOLOGUES-EST was benchmarked with genomes and transcriptomes of Arabidopsis thaliana and Hordeum vulgare, available at http:https://floresta.eead.csic.es/plant-pan-genomes, and used to analyze the pan-genomes of Brachypodium distachyon and Brachypodium hybridum (press release).
Two tutorials are available:
-
Pangenome analysis of plant transcripts and coding sequences, published in 2022.
-
From genomes to pangenomes: understanding variation among individuals and species, which includes step by step instructions for both bacterial and plant data, first released in 2017.
Installation instructions, including the bioconda package, are available in the manual and the README.txt file.
| version | HTML |
|---|---|
| original, for the analysis of bacterial pan-genomes | manual |
| EST, for the analysis of intra-species eukaryotic pan-genomes, tested on plants | manual-est |
A Docker image is also available with GET_HOMOLOGUES bundled with GET_PHYLOMARKERS, ready to use. The GET_PHYLOMARKERS manual explains how to use nucleotide & peptide clusters produced by GET_HOMOLOGUES to compute robust multi-gene and pangenome phylogenies.
A related piece of software was released in 2023 called GET_PANGENES, which takes FASTA and GFF files as input and explicitely considers gene collinearity.
The code is regularly patched (see CHANGES.txt in each release, and has been used in a variety of studies (see citing papers here and here, respectively).
We kindly ask you to report errors or bugs to the authors and to acknowledge the use of the software in scientific publications.
GET_HOMOLOGUES is part of the INB/ELIXIR-ES resources portfolio:
Funding: Fundacion ARAID, Consejo Superior de Investigaciones Cientificas, DGAPA-PAPIIT UNAM, CONACyT, FEDER, MINECO, DGA-Obra Social La Caixa.