This repository includes the source code and test data for the paper https://arxiv.org/abs/2401.05638
We are continuously improving the code and the documentation. If you have any questions or suggestions, please feel free to contact us.
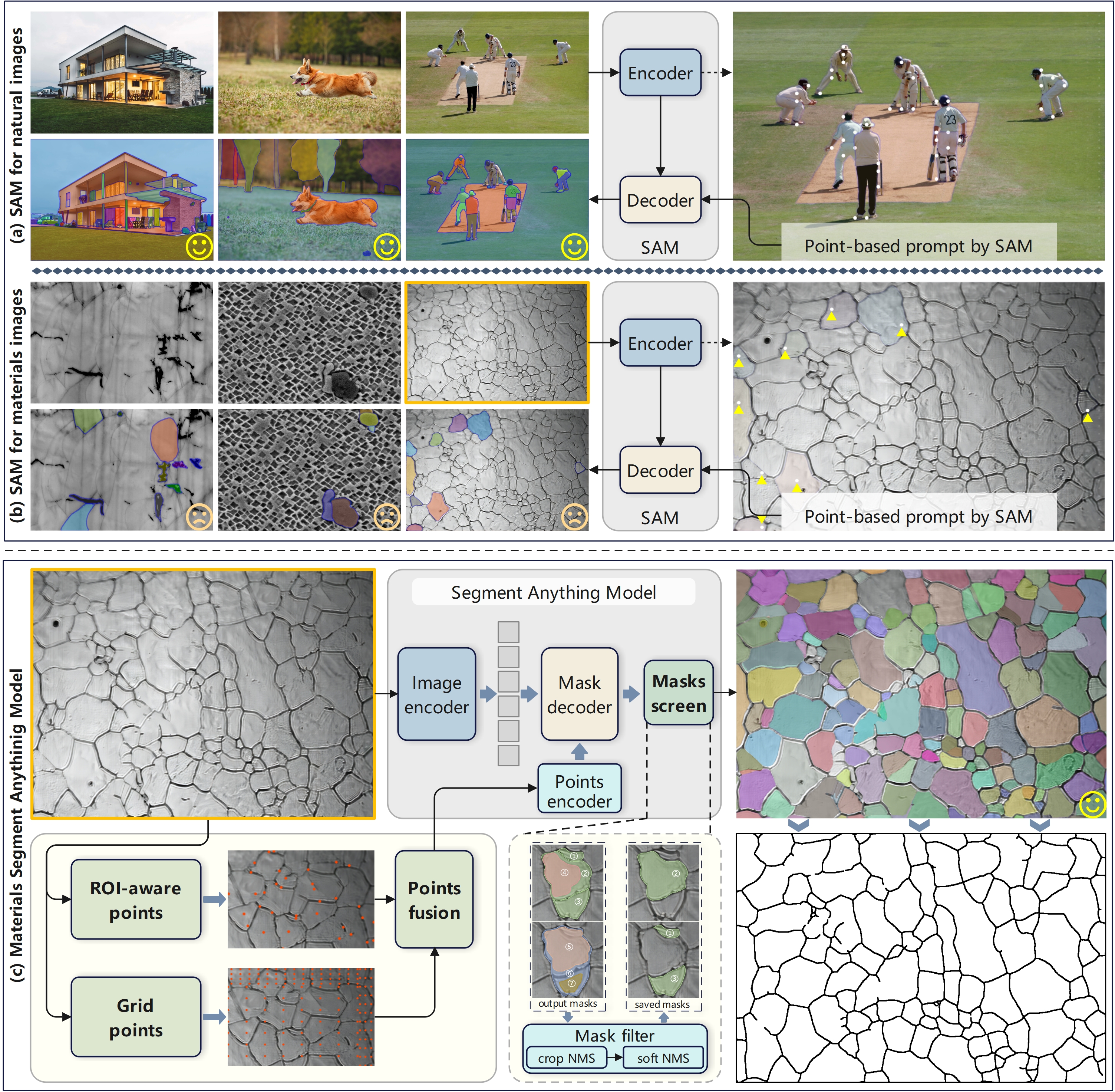
Efficient and accurate extraction of microstructures in micrographs of materials is essential in process optimization and the exploration of structure-property relationships. Deep learning-based image segmentation techniques that rely on manual annotation are laborious and time-consuming and hardly meet the demand for model transferability and generalization on various source images. Segment Anything Model (SAM), a large visual model with powerful deep feature representation and zero-shot generalization capabilities, has provided new solutions for image segmentation. However, directly applying SAM to segmenting microstructures in microscopy images without human annotation cannot achieve the expected results, as the difficulty of adapting its native prompt engineering to the dense and dispersed characteristics of key microstructures in different materials. In this paper, we propose MatSAM, a general and efficient microstructure extraction solution based on SAM. A simple yet effective point-based prompt generation strategy is designed, grounded on the distribution and shape of microstructures. Specifically, in an unsupervised and training-free way, it adaptively generates prompt points for different microscopy images, fuses the centroid points of the coarsely extracted region of interest (ROI) and native grid points, and integrates corresponding post-processing operations for quantitative characterization of microstructures of materials. For common microstructures including grain boundary and multiple phases, MatSAM achieves superior zero-shot segmentation performance to conventional rule-based methods and is even preferable to supervised learning methods evaluated on 16 microscopy datasets whose micrographs are imaged by the optical microscope (OM) and scanning electron microscope (SEM). Especially, on 4 public datasets, MatSAM shows unexpected competitive segmentation performance against their specialist models. We believe that, without the need for human labeling, MatSAM can significantly reduce the cost of quantitative characterization and statistical analysis of extensive microstructures of materials, and thus accelerate the design of new materials.
We suggest using conda to create a virtual environment listed in /requirements.txt.
The required package gala needs manual installation.
For ease of reproducibility, we have provided the source code in the form of a jupyter notebook. The notebook is divided into sections, each of which corresponds to a step in the proposed method. Sets of hyperparameters are provided in the notebook, and the user can adjust them according to the specific requirements of the dataset.
-
/notebook/matsam_example.ipynb gives an example of how to use the code to extract grains from a polycrystalline microscopy image.
-
/notebook/matsam_public_datasets.ipynb gives the evaluation results of the proposed method on the public datasets:
- NBS-2 (Ni-superalloys-Super2 in https://doi.org/10.1038/s41524-022-00878-5)
- NBS-3 (Ni-superalloys-Super4 in https://doi.org/10.1038/s41524-022-00878-5)
- AZA (XCT-Al-Zn-alloy in https://doi.org/10.1016/j.matchar.2020.110119)
- UHCS (Ultrahigh-carbon-steel in https://doi.org/10.1016/j.actamat.2023.119086)
-
The checkpoints can be downloaded by the links:
- ViT-H(default): ViT-H SAM model
- ViT-L: ViT-L SAM model
- ViT-B: ViT-B SAM model
The downloaded checkpoints should be placed in directory /checkpoints/.
For in-house datasets, we will provide partial test data and the corresponding ground truth in the future updates.