Link to Journal of Dairy Science
You will need to get access key via Miel Hostens ([email protected])
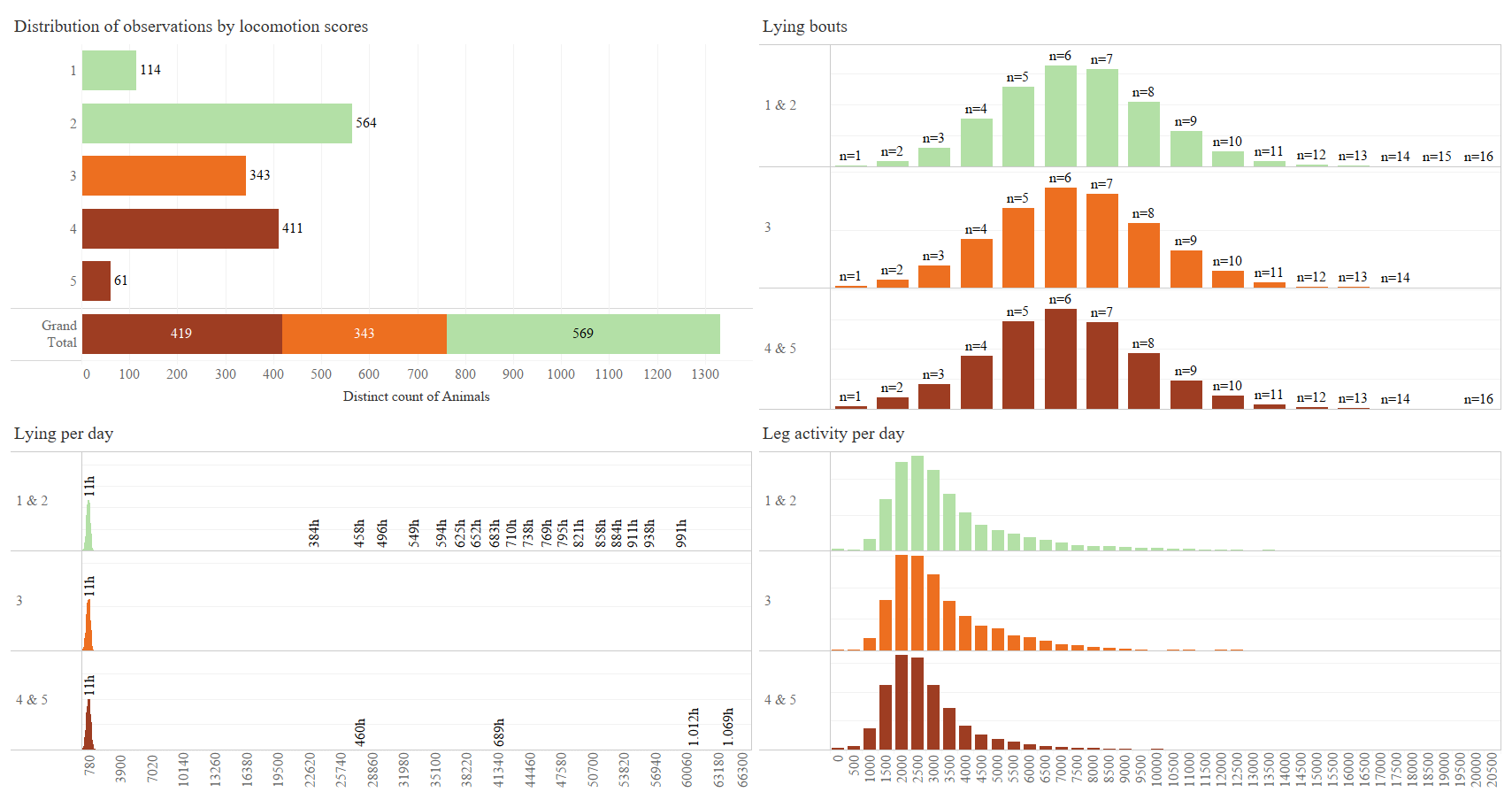
The data contains data from 8 farms using NEDAP sensors. An observation from the NEDAP sensores was collected together with locomotion scores from the animals.
- HerdIdentifier - Identifier for the herd
- Animalidentifier : Identifier for the animal (her eartag)
- AnimalNumber : Secondary identifier for the animal
- CalvingTime : Date of calving
- SensorDaysInMilk : Days in milk at sensor observation
- SensorDate : Date at sensor observation
- SensorType: Multiple sensor observations were obtained, the type identifies which ones are usefull
- SensorValue: Sensor value
- ObservationDate: Date at which the locomotion or bcs scores were observed
- ScoreDaysInMilk : Days in milk at which the locomotion or bcs scores were observed
- ScoreType : Locomotion or BCS score
- ScoreValue : Score value
- ObservationDays : Days from sensor to score observation moment
The underlying images can be seen in the
:
- Make sure to have the data downloaded on disk
- Make sure to have the data referenced correctly in the workbook
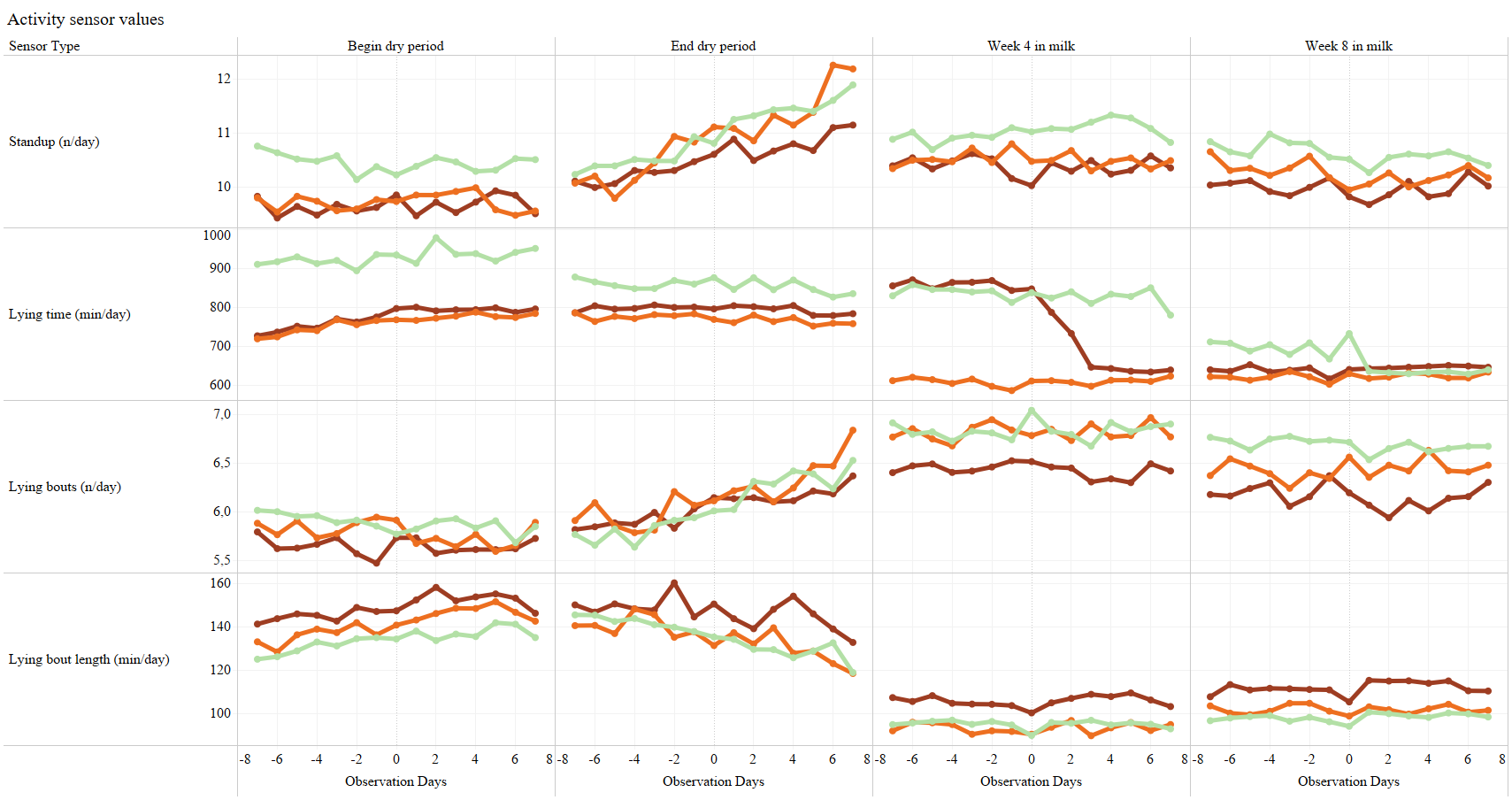
- Link to model Standups
- Link to model Leg Activity
- Link to model Lying Time
- Link to model Lying Bouts
- Link to model Lying Bout Length
- error bar -> CI DONE
- -1 tot +1 DONE
- referentie tijdstip naar 4w DONE
- lying bouts contains DataType DONE
- Add fixed effect pre and post partum DONE
- heifers separate analysis –> parity 2, 3 and 4+ depending on group size DONE
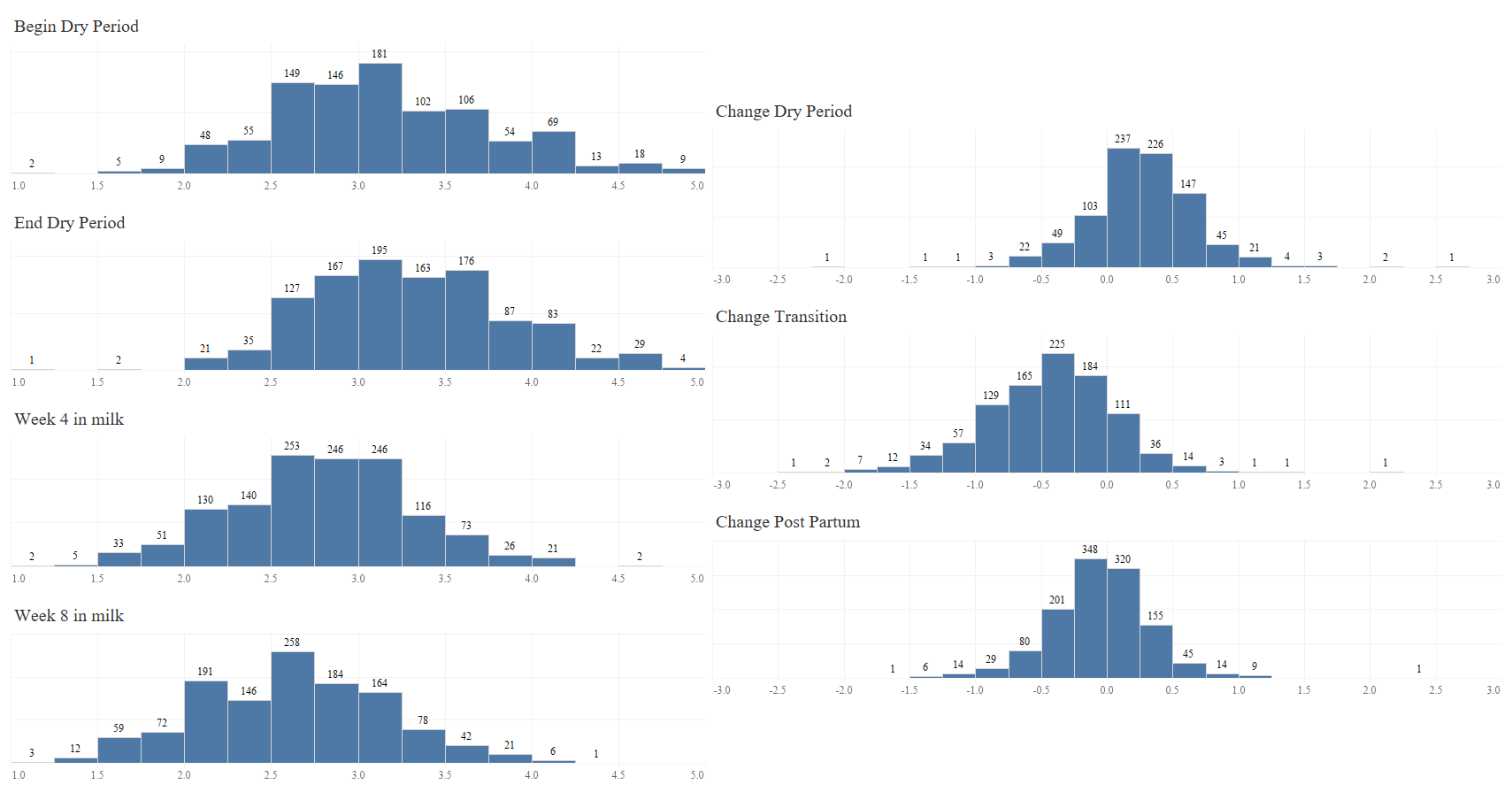
- grouping loco scores: 1&2, 3, 4&5 DONE
- loco score 3 as reference, 1&2 as reference as well. check both for contrast (relevel) DONE
- QQ plot log link DONE
- partum x ls x moment DONE BUT UNABLE TO FIT
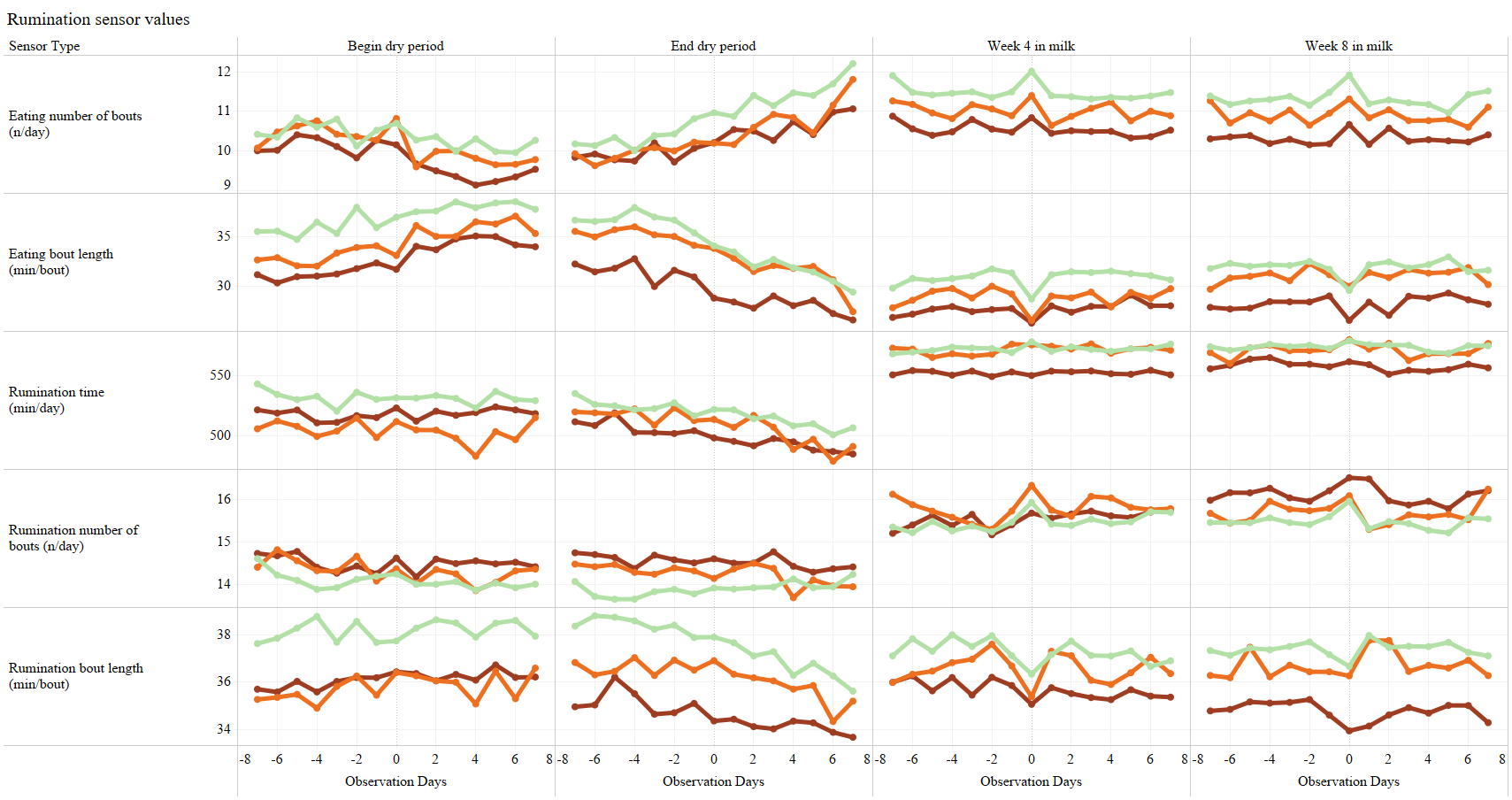
- types of behavior for analysis: eating time/day, rumination time/day, eating+rumination time per day, lying time per day, number of steps per day (legactivity)
- discussion: correlation between behavior of residuals in neck & leg sensor data output
- discussion: visualisation
- after definitive results: involve Klaas Frankena
- Wald -> profile likelihood
- Force model to be fact1*fact2 -> fact1:fact2 + fact2 (this is factor one nested within factor 2)
- Short values of lyingboutlength can be the cutoff on 24h DONE
- cutoff to 1440 DONE
- Use log of lyingboutlength DONE
- LyingBout only -1 ??? - if no diff leave, if yes diff keep that model DONE (See Lying Bouts folder
- At model scale, a difference was detected … is good way of writing the article
- When comparing models, keep ALL main effects in, when CONFINT then drop DONE
- Request lying bout and lying bout length
- Check LSM and Figure LSM output (DONE)
- Check CI instead of SE (DONE)
- Percentage calculations tableau
- Changed trend symbol in figure
- Increase from -1 and 1 to -2 and 2 window around scoring
- Added need for 4 sensorvalues per average
- Add dplyr for distinct numbers
- Added calving season
- Added association models