This repository reproduces results from the paper Personalizing renal replacement therapy initiation in the intensive care unit: a reinforcement learning-based strategy with external validation on the AKIKI randomized controlled trials [medRxiv].
This repository is written and maintained by François Grolleau ([email protected]).
- The Learning folder includes code related to the section titled “Learning an optimal strategy” in the paper.
It contains the following files.
dataprep_dev.R details the preprocessing and multiple imputations of the MIMIC data.
data_description_dev.R reproduces Table 1 and Figure S3 (A) for the MIMIC.
dWOLS.R details the dWOLS estimation procedure for optimal strategy learning in the MIMIC data.
figS2_A.R reproduces Figure S2 (A) for MIMIC.
figS2_B.R reproduces Figure S2 (B) for MIMIC.
functions.R contains handy functions defactorize and rubinr to “defactorize” dataframes and compute Rubin's rule respectively.
export_feather.R export the imputed development dataframes as feather files for downstream use in Python.
crossfit_ADR_eval.py details the cross-fitted advantage doubly robust estimator for strategy evaluation for hospital-free days at day 60, using the MIMIC data (apparent effect).
- The Evaluation folder includes code related to the section titled “External validation” in the paper.
It contains the following files.
akiki_1_2_preprocessing.R details the preprocessing and multiple imputations of the AKIKI trial data.
data_description_val.R reproduces Table 1 and Figure S3 (B) for the AKIKI trial data.
get_predictions_val.R get recommendations from the “crude” and “stringent” learned strategies, and export the imputed validation dataframes as feather files for downstream use in Python.
n_rrt_eval.py details importance sampling for policy evaluation for the proportion of patients who would initiate renal replacement therapy within three days, using the AKIKI trial data.
hmor_eval.py details importance sampling for policy evaluation for hospital mortality, using the AKIKI trial data. This file is adapted from n_rrt_eval.py.
crossfit_ADR_eval.py details the cross-fitted advantage doubly robust estimator for strategy evaluation with terminal states for hospital-free days at day 60, using the AKIKI trial data.
fig2.R reproduces Figure 2 of the paper.
fig3_A.R reproduces Figure 3 (A) of the paper.
fig3_B.R reproduces Figure 3 (A) of the paper.
fig4.R reproduces Figure 2 of the paper.
- The Shiny folder includes the code producing the webapp for our dynamic decision support system.
It contains the following files and folders.
ui.R for the user interface shiny file.
server.R for the server shiny file.
www a folder that contains the cress.png and uparis.png logos.
img a folder that contains screenshots from our webapp.
Sankey diagrams were produced with the files aforementioned R scripts corresponding to Figure 3 and S2. The HTML files produced were then imported as resolution-independent images with SVG Crowbar and colored in Adobe Illustrator.
Bootstrapping was performed with the Speedboot Python library that we developed during the course of this study. The library is available on Pypi with documentation accessible here.
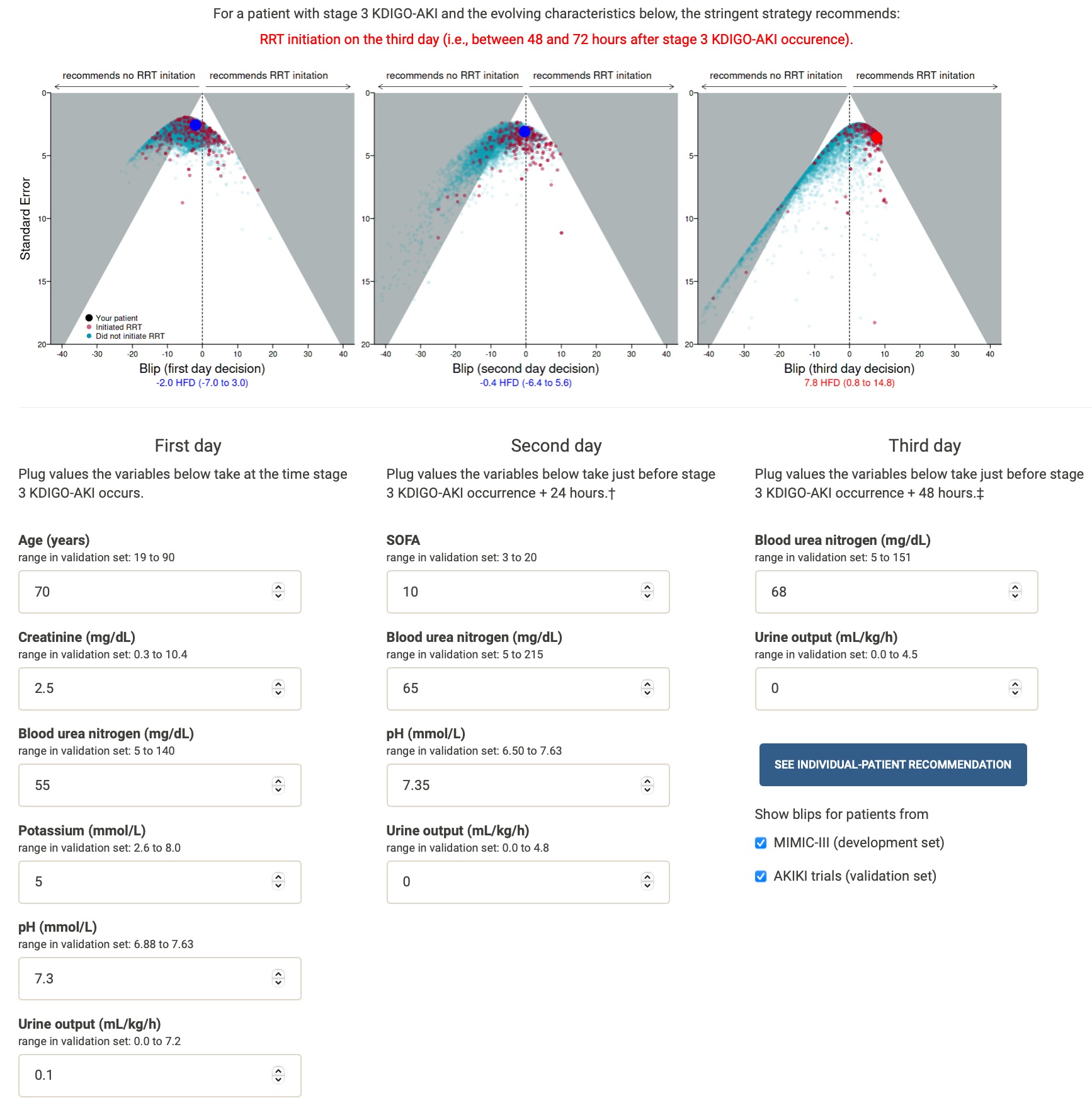
Available at http:https://dynamic-rrt.eu/.
This applied work heavily relies on the following two technical papers:
-
Wallace MP and Moodie EEM. Doubly-robust dynamic treatment regimen estimation via weighted least squares. Biometrics 2015; 71: 636–44.
-
Nie X, Brunskill E, and Wager S. Learning when-to-treat policies. Journal of the American Statistical Association 2021; 116: 392–409. [arXiv].