The repository is an official implementation of SaProt: Protein Language Modeling with Structure-aware Vocabulary.
If you have any question about the paper or the code, feel free to raise an issue! Saprot should outperform ESM-2 in most tasks under fair evaluation settings.
The laboratory is hiring research assistants, interns, doctoral students, and postdoctoral researchers. Please contact the corresponding author for details.
实验室招聘科研助理,实习生,博士生和博士后,请联系通讯作者
Table of contents
- 2024/08/14: over 20 outstanding researchers in Biology&Bioinformatics have joined SaprotHub as co-authors. Joining us and contribute.
- 2024/05/13: We developed SaprotHub to make protein language model training accessible to all biologists. Go.
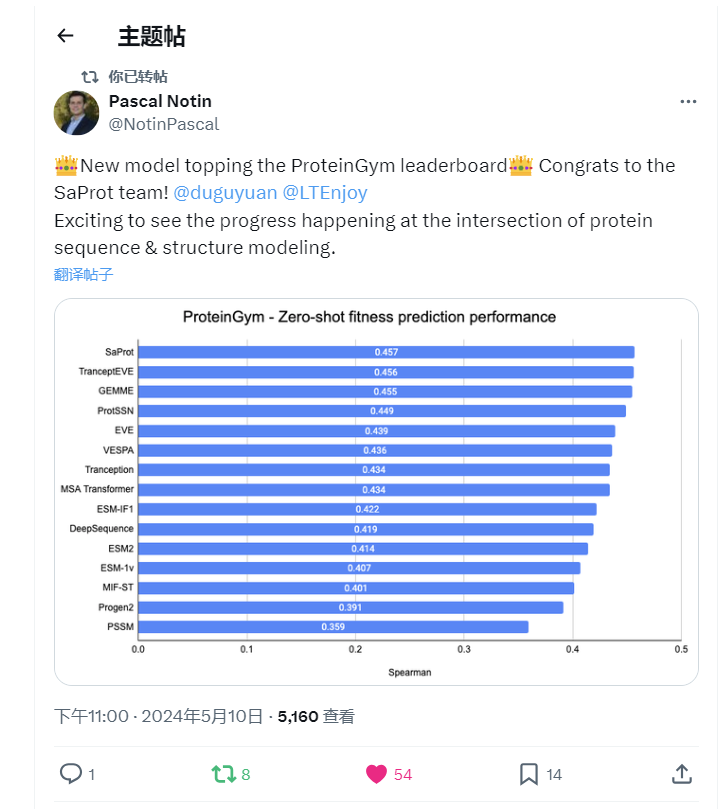
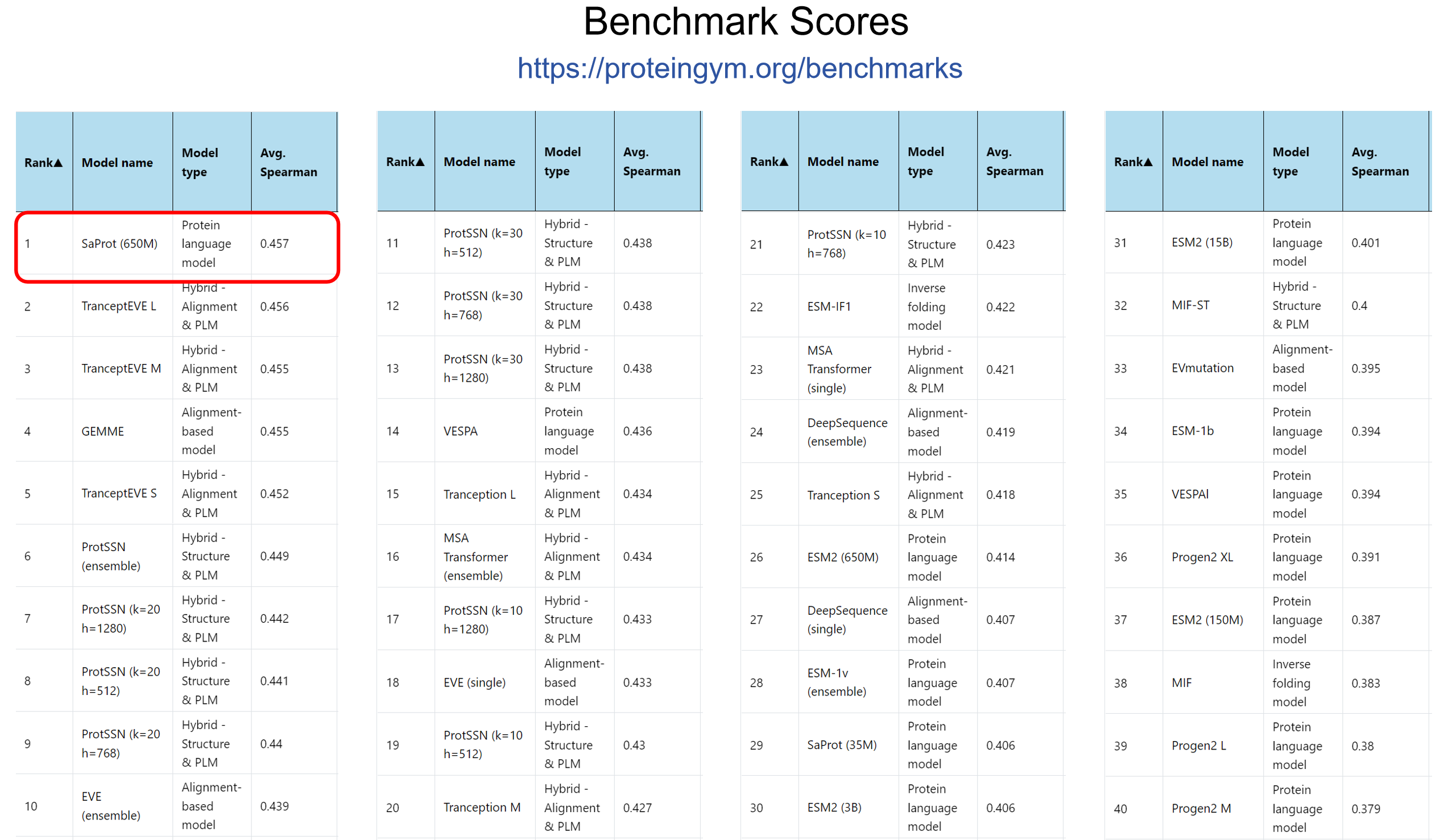
- 2024/05/13: SaProt ranked #1st on the public ProteinGym benchmark in April2024, while other top-ranked models are hybrid and mutation-specialized model.🎉🎉🎉! See here.
- 2024/04/18: We found a slight difference for EC and GO evaluation and updated the re-evaluated results (see issue #23 for details).
- 2024/03/08: We uploaded a simple function to make zero-shot prediction of mutational effect (see example below).
- 2024/01/17: Our paper has been accepted as ICLR 2024 spotlight 🎉🎉🎉!
- 2023/10/30: We release a pre-trained SaProt 35M model and a 35M residue-sequence-only version of SaProt (for comparison)! The residue-sequence-only SaProt (without 3Di token) performs highly similar to the official ESM-2 35M model. (see Results below).
- 2023/10/30: We released the results by using ESMFold structures. See Table below
We propose a structure-aware vocabulary for protein language modeling. The vocabulary is constructed by encoding the
protein structure into discrete 3D tokens by using the foldseek. We combine the residue tokens and the structure tokens to form a structure-aware sequence.
Through large-scale pre-training, our model, i.e. SaProt, can learn the relationship between the structure and the sequence.
For more details, please refer to our paper https://www.biorxiv.org/content/10.1101/2023.10.01.560349v2.

conda create -n SaProt python=3.10
conda activate SaProt
bash environment.sh
We provide two ways to use SaProt, including through huggingface class and through the same way in esm github. Users can choose either one to use.
| Name | Size | Dataset |
|---|---|---|
| SaProt_35M_AF2 | 35M parameters | 40M AF2 structures |
| SaProt_650M_PDB | 650M parameters | 40M AF2 structures (phase1) + 60K PDB structures (phase2) |
| SaProt_650M_AF2 | 650M parameters | 40M AF2 structures |
Some experimental results are listed below. For more details, please refer to our paper.
| Model | ClinVar | ProteinGym | Thermostability | HumanPPI | Metal Ion Binding | EC | GO-MF | GO-BP | GO-CC | DeepLoc-Subcellular | DeepLoc-Binary |
|---|---|---|---|---|---|---|---|---|---|---|---|
| AUC | Spearman's ρ | Spearman's ρ | Acc% | Acc% | Fmax | Fmax | Fmax | Fmax | Acc% | Acc% | |
| ESM-2 (35M) | 0.722 | 0.339 | 0.669 | 80.79 | 73.08 | 0.825 | 0.616 | 0.416 | 0.404 | 76.58 | 91.60 |
| SaProt-Seq (35M) | 0.738 | 0.337 | 0.672 | 80.56 | 73.23 | 0.821 | 0.608 | 0.413 | 0.403 | 76.67 | 91.16 |
| SaProt (35M) | 0.794 | 0.392 | 0.692 | 81.11 | 74.29 | 0.847 | 0.642 | 0.431 | 0.418 | 78.09 | 91.97 |
| Model | ClinVar | ProteinGym | Thermostability | HumanPPI | Metal Ion Binding | EC | GO-MF | GO-BP | GO-CC | DeepLoc-Subcellular | DeepLoc-Binary |
|---|---|---|---|---|---|---|---|---|---|---|---|
| AUC | Spearman's ρ | Spearman's ρ | Acc% | Acc% | Fmax | Fmax | Fmax | Fmax | Acc% | Acc% | |
| ESM-2 (650M) | 0.862 | 0.475 | 0.680 | 76.67 | 71.56 | 0.868 | 0.670 | 0.473 | 0.470 | 82.09 | 91.96 |
| SaProt (650M) | 0.909 | 0.478 | 0.724 | 86.41 | 75.75 | 0.882 | 0.682 | 0.486 | 0.479 | 85.57 | 93.55 |
We compare structures predicted by AF2 or ESMFold, which is shown below:
| model | ClinVar | ProteinGym | Thermostability | HumanPPI | Metal Ion Binding | EC | GO-MF | GO-BP | GO-CC | DeepLoc-Subcellular | DeepLoc-Binary |
|---|---|---|---|---|---|---|---|---|---|---|---|
| AUC | Spearman's ρ | Spearman's ρ | Acc% | Acc% | Fmax | Fmax | Fmax | Fmax | Acc% | Acc% | |
| SaProt (ESMFold) | 0.896 | 0.455 | 0.717 | 85.78 | 74.10 | 0.871 | 0.678 | 0.480 | 0.474 | 82.82 | 93.19 |
| SaProt (AF2) | 0.909 | 0.478 | 0.724 | 86.41 | 75.75 | 0.882 | 0.682 | 0.486 | 0.479 | 85.57 | 93.55 |
SaProt achieved first position on ProteinGym benchmark! The checkpoint was trained on Sep. 2023.
The following code shows how to load the model based on huggingface class. Note masking lower pLDDT regions for AF2 structures is beneficial ,see below.
from transformers import EsmTokenizer, EsmForMaskedLM
model_path = "/your/path/to/SaProt_650M_AF2" # Note this is the directory path of SaProt, not the ".pt" file
tokenizer = EsmTokenizer.from_pretrained(model_path)
model = EsmForMaskedLM.from_pretrained(model_path)
#################### Example ####################
device = "cuda"
model.to(device)
seq = "M#EvVpQpL#VyQdYaKv" # Here "#" represents lower plDDT regions (plddt < 70)
tokens = tokenizer.tokenize(seq)
print(tokens)
inputs = tokenizer(seq, return_tensors="pt")
inputs = {k: v.to(device) for k, v in inputs.items()}
outputs = model(**inputs)
print(outputs.logits.shape)
"""
['M#', 'Ev', 'Vp', 'Qp', 'L#', 'Vy', 'Qd', 'Ya', 'Kv']
torch.Size([1, 11, 446])
"""
User could also load SaProt by esm implementation. The checkpoint is
stored in the same huggingface folder, named SaProt_650M_AF2.pt. We provide a function to load the model.
from utils.esm_loader import load_esm_saprot
model_path = "/your/path/to/SaProt_650M_AF2.pt"
model, alphabet = load_esm_saprot(model_path)
We provide a function to convert a protein structure into a structure-aware sequence. The function calls the
foldseek
binary file to encode the structure. You can download the binary file from here and place it in the bin folder
. The following code shows how to use it.
from utils.foldseek_util import get_struc_seq
pdb_path = "example/8ac8.cif"
# Extract the "A" chain from the pdb file and encode it into a struc_seq
# pLDDT is used to mask low-confidence regions if "plddt_mask" is True. Please set it to True when
# use AF2 structures for best performance.
parsed_seqs = get_struc_seq("bin/foldseek", pdb_path, ["A"], plddt_mask=False)["A"]
seq, foldseek_seq, combined_seq = parsed_seqs
print(f"seq: {seq}")
print(f"foldseek_seq: {foldseek_seq}")
print(f"combined_seq: {combined_seq}")
We provide a function to predict the mutational effect of a protein sequence. The example below shows how to predict the mutational effect at a specific position. If using the AF2 structure, we strongly recommend that you add pLDDT mask (see below).
from model.saprot.saprot_foldseek_mutation_model import SaprotFoldseekMutationModel
config = {
"foldseek_path": None,
"config_path": "/your/path/to/SaProt_650M_AF2", # Note this is the directory path of SaProt, not the ".pt" file
"load_pretrained": True,
}
model = SaprotFoldseekMutationModel(**config)
tokenizer = model.tokenizer
device = "cuda"
model.eval()
model.to(device)
seq = "M#EvVpQpL#VyQdYaKv" # Here "#" represents lower plDDT regions (plddt < 70)
# Predict the effect of mutating the 3rd amino acid to A
mut_info = "V3A"
mut_value = model.predict_mut(seq, mut_info)
print(mut_value)
# Predict all effects of mutations at 3rd position
mut_pos = 3
mut_dict = model.predict_pos_mut(seq, mut_pos)
print(mut_dict)
# Predict probabilities of all amino acids at 3rd position
mut_pos = 3
mut_dict = model.predict_pos_prob(seq, mut_pos)
print(mut_dict)
"""
0.7908501625061035
{'V3A': 0.7908501625061035, 'V3C': -0.9117952585220337, 'V3D': 2.7700226306915283, 'V3E': 2.3255627155303955, 'V3F': 0.2094242423772812, 'V3G': 2.699633836746216, 'V3H': 1.240191102027893, 'V3I': 0.10231903940439224, 'V3K': 1.804598093032837,
'V3L': 1.3324960470199585, 'V3M': -0.18938277661800385, 'V3N': 2.8249857425689697, 'V3P': 0.40185314416885376, 'V3Q': 1.8361762762069702, 'V3R': 1.1899691820144653, 'V3S': 2.2159857749938965, 'V3T': 0.8813426494598389, 'V3V': 0.0, 'V3W': 0.5853186249732971, 'V3Y': 0.17449656128883362}
{'A': 0.021275954321026802, 'C': 0.0038764977362006903, 'D': 0.15396881103515625, 'E': 0.0987202599644661, 'F': 0.011895398609340191, 'G': 0.14350374042987823, 'H': 0.03334535285830498, 'I': 0.010687196627259254, 'K': 0.058634623885154724, 'L': 0.03656982257962227, 'M': 0.00798324216157198, 'N': 0.16266827285289764, 'P': 0.014419485814869404, 'Q': 0.06051575019955635, 'R': 0.03171204403042793, 'S': 0.08847439289093018, 'T': 0.023291070014238358, 'V': 0.009647775441408157, 'W': 0.017323188483715057, 'Y': 0.011487090960144997}
"""
We provide the dataset for pre-training SaProt. The dataset can be downloaded from here.
We provide datasets that are used in the paper. Datasets can be downloaded from here.
Once downloaded, the datasets need to be decompressed and placed in the LMDB folder for supervised fine-tuning.
We provide a script to fine-tune SaProt on the datasets. The following code shows how to fine-tune SaProt on specific
downstream tasks. Before running the code, please make sure that the datasets are placed in the LMDB folder and the
huggingface version of SaProt 650M model is placed in the weights/PLMs folder. Note that the default training setting is not as
same as in the paper because of the hardware limitation for different users. We recommend users to modify the yaml file
flexibly based on their own conditions (i.e. batch_size, devices and accumulate_grad_batches).
# Fine-tune SaProt on the Thermostability task
python scripts/training.py -c config/Thermostability/saprot.yaml
# Fine-tune ESM-2 on the Thermostability task
python scripts/training.py -c config/Thermostability/esm2.yaml
If you want to record the training process using wandb, you could modify the config file and set Trainer.logger = True
and then paste your wandb API key in the config key setting.os_environ.WANDB_API_KEY.
We provide a script to evaluate the zero-shot performance of models (foldseek binary file is required to be placed in
the bin folder):
# Evaluate the zero-shot performance of SaProt on the ProteinGym benchmark
python scripts/mutation_zeroshot.py -c config/ProteinGym/saprot.yaml
# Evaluate the zero-shot performance of ESM-2 on the ProteinGym benchmark
python scripts/mutation_zeroshot.py -c config/ProteinGym/esm2.yaml
The results will be saved in the output/ProteinGym folder.
For ClinVar benchmark, you can use the following script to calculate the AUC metric:
# Evaluate the zero-shot performance of SaProt on the ClinVar benchmark
python scripts/mutation_zeroshot.py -c config/ClinVar/saprot.yaml
python scripts/compute_clinvar_auc.py -c config/ClinVar/saprot.yaml
If you find this repository useful, please cite our paper:
@article{su2023saprot,
title={SaProt: Protein Language Modeling with Structure-aware Vocabulary},
author={Su, Jin and Han, Chenchen and Zhou, Yuyang and Shan, Junjie and Zhou, Xibin and Yuan, Fajie},
journal={bioRxiv},
year={2023},
publisher={Cold Spring Harbor Laboratory}